DE genes between T cell groups
Mechthild Lütge
Dec 2022
Last updated: 2022-12-12
Checks: 6 1
Knit directory: humanCardiacFibroblasts/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210903) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 61f9e3b. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: data/GSEA/
Ignored: data/humanFibroblast/
Ignored: figure/DEgenesGZplusSG_Groups.Rmd/.DS_Store
Unstaged changes:
Modified: analysis/projectSignatures.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/projectSignatures.Rmd) and
HTML (docs/projectSignatures.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 307a2ca | mluetge | 2022-12-12 | update pat ID SG29 |
| html | 307a2ca | mluetge | 2022-12-12 | update pat ID SG29 |
| Rmd | 66c2208 | mluetge | 2022-12-08 | compare T cell groups and project gene signatures |
| html | 66c2208 | mluetge | 2022-12-08 | compare T cell groups and project gene signatures |
load packages
suppressPackageStartupMessages({
library(SingleCellExperiment)
library(tidyverse)
library(Seurat)
library(magrittr)
library(dplyr)
library(purrr)
library(ggplot2)
library(here)
library(runSeurat3)
library(ggsci)
library(pheatmap)
library(ggpubr)
library(RColorBrewer)
library(viridis)
})heatmap function
avgHeatmap <- function(seurat, selGenes, colVecIdent, colVecCond=NULL,
ordVec=NULL, gapVecR=NULL, gapVecC=NULL,cc=FALSE,
cr=FALSE, condCol=FALSE){
selGenes <- selGenes$gene
## assay data
clusterAssigned <- as.data.frame(Idents(seurat)) %>%
dplyr::mutate(cell=rownames(.))
colnames(clusterAssigned)[1] <- "ident"
seuratDat <- GetAssayData(seurat)
## genes of interest
genes <- data.frame(gene=rownames(seurat)) %>%
mutate(geneID=gsub("^.*\\.", "", gene)) %>% filter(geneID %in% selGenes)
## matrix with averaged cnts per ident
logNormExpres <- as.data.frame(t(as.matrix(
seuratDat[which(rownames(seuratDat) %in% genes$gene),])))
logNormExpres <- logNormExpres %>% dplyr::mutate(cell=rownames(.)) %>%
dplyr::left_join(.,clusterAssigned, by=c("cell")) %>%
dplyr::select(-cell) %>% dplyr::group_by(ident) %>%
dplyr::summarise_all(mean)
logNormExpresMa <- logNormExpres %>% dplyr::select(-ident) %>% as.matrix()
rownames(logNormExpresMa) <- logNormExpres$ident
logNormExpresMa <- t(logNormExpresMa)
rownames(logNormExpresMa) <- gsub("^.*?\\.","",rownames(logNormExpresMa))
## remove genes if they are all the same in all groups
ind <- apply(logNormExpresMa, 1, sd) == 0
logNormExpresMa <- logNormExpresMa[!ind,]
genes <- genes[!ind,]
## color columns according to cluster
annotation_col <- as.data.frame(gsub("(^.*?_)","",
colnames(logNormExpresMa)))%>%
dplyr::mutate(celltype=gsub("(_.*$)","",colnames(logNormExpresMa)))
colnames(annotation_col)[1] <- "col1"
annotation_col <- annotation_col %>%
dplyr::mutate(cond = gsub(".*_","",col1)) %>%
dplyr::select(cond, celltype)
rownames(annotation_col) <- colnames(logNormExpresMa)
ann_colors = list(
cond = colVecCond,
celltype=colVecIdent)
if(is.null(ann_colors$cond)){
annotation_col$cond <- NULL
}
## adjust order
logNormExpresMa <- logNormExpresMa[selGenes,]
if(is.null(ordVec)){
ordVec <- levels(seurat)
}
logNormExpresMa <- logNormExpresMa[,ordVec]
## scaled row-wise
pheatmap(logNormExpresMa, scale="row" ,treeheight_row = 0, cluster_rows = cr,
cluster_cols = cc, border_color = NA,
color = colorRampPalette(c("#2166AC", "#F7F7F7", "#B2182B"))(50),
annotation_col = annotation_col, cellwidth=15, cellheight=10,
annotation_colors = ann_colors, gaps_row = gapVecR, gaps_col = gapVecC)
}sign plot funct
## adapted from CellMixS
visGroup_adapt <- function (sce,group,dim_red = "TSNE",col_group=pal_nejm()(8))
{
if (!is(sce, "SingleCellExperiment")) {
stop("Error:'sce' must be a 'SingleCellExperiment' object.")
}
if (!group %in% names(colData(sce))) {
stop("Error: 'group' variable must be in 'colData(sce)'")
}
cell_names <- colnames(sce)
if (!dim_red %in% "TSNE") {
if (!dim_red %in% reducedDimNames(sce)) {
stop("Please provide a dim_red method listed in reducedDims of sce")
}
red_dim <- as.data.frame(reducedDim(sce, dim_red))
}
else {
if (!"TSNE" %in% reducedDimNames(sce)) {
if ("logcounts" %in% names(assays(sce))) {
sce <- runTSNE(sce)
}
else {
sce <- runTSNE(sce, exprs_values = "counts")
}
}
red_dim <- as.data.frame(reducedDim(sce, "TSNE"))
}
colnames(red_dim) <- c("red_dim1", "red_dim2")
df <- data.frame(sample_id = cell_names, group_var = colData(sce)[,
group], red_Dim1 = red_dim$red_dim1, red_Dim2 = red_dim$red_dim2)
t <- ggplot(df, aes_string(x = "red_Dim1", y = "red_Dim2")) +
xlab(paste0(dim_red, "_1")) + ylab(paste0(dim_red, "_2")) +
theme_void() + theme(aspect.ratio = 1,
panel.grid.minor = element_blank(),
panel.grid.major = element_line(color = "grey", size = 0.3))
t_group <- t + geom_point(size = 1.5, alpha = 0.8,
aes_string(color = "group_var")) +
guides(color = guide_legend(override.aes = list(size = 1),
title = group)) + ggtitle(group)
if (is.numeric(df$group_var)) {
t_group <- t_group + scale_color_viridis(option = "D")
}
else {
t_group <- t_group + scale_color_manual(values = col_group)
}
t_group
}set dir
basedir <- here()
seurat <- readRDS(file = paste0(basedir,
"/data/humanHeartsPlusGraz_intPatients_merged",
"labeled_woHH_seurat.rds"))
Idents(seurat) <- seurat$integrated_snn_res.0.6color vectors
colPal <- c(pal_igv()(12),
pal_aaas()(10))[1:length(levels(seurat))]
colTec <- pal_jama()(length(unique(seurat$technique)))
colSmp <- c(pal_uchicago()(9), pal_npg()(10), pal_aaas()(10),
pal_jama()(7))[1:length(unique(seurat$dataset))]
colCond <- pal_npg()(length(unique(seurat$cond)))
colID <- c(pal_jco()(10), pal_npg()(10), pal_futurama()(10),
pal_d3()(10))[1:length(unique(seurat$ID))]
colOrig <- pal_aaas()(length(unique(seurat$origin)))
colIso <- pal_nejm()(length(unique(seurat$isolation)))
colProc <- pal_aaas()(length(unique(seurat$processing)))
colTgrp <- c("#d70700", "#00239a", "#1f7a1f")
colLab <- c("#c08b65", "#ba4e45", "#d4cc84", "#546f82", "#5c5cdf",
"#80396e", "#8d5639", "#779462", "#800000FF", "#d87c15",
"#FFA319FF", "#FF95A8FF")
names(colLab) <- c("EndoEC", "Tcell","resMacrophage", "Fibroblast",
"infMacrophage", "Perivascular","Cardiomyocyte",
"Endothelial","Adipocytes","NeuralCells","SMC","LEC")
names(colTgrp) <- c("TcellHigh", "TcellLow", "TcellInt" )
names(colPal) <- levels(seurat)
names(colTec) <- unique(seurat$technique)
names(colSmp) <- unique(seurat$dataset)
names(colCond) <- unique(seurat$cond)
names(colID) <- unique(seurat$ID)
names(colOrig) <- unique(seurat$origin)
names(colIso) <- unique(seurat$isolation)
names(colProc) <- unique(seurat$processing)vis data
clusters
DimPlot(seurat, reduction = "umap", cols=colPal)+
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
panel.grid.minor = element_blank()) +
xlab("UMAP1") +
ylab("UMAP2")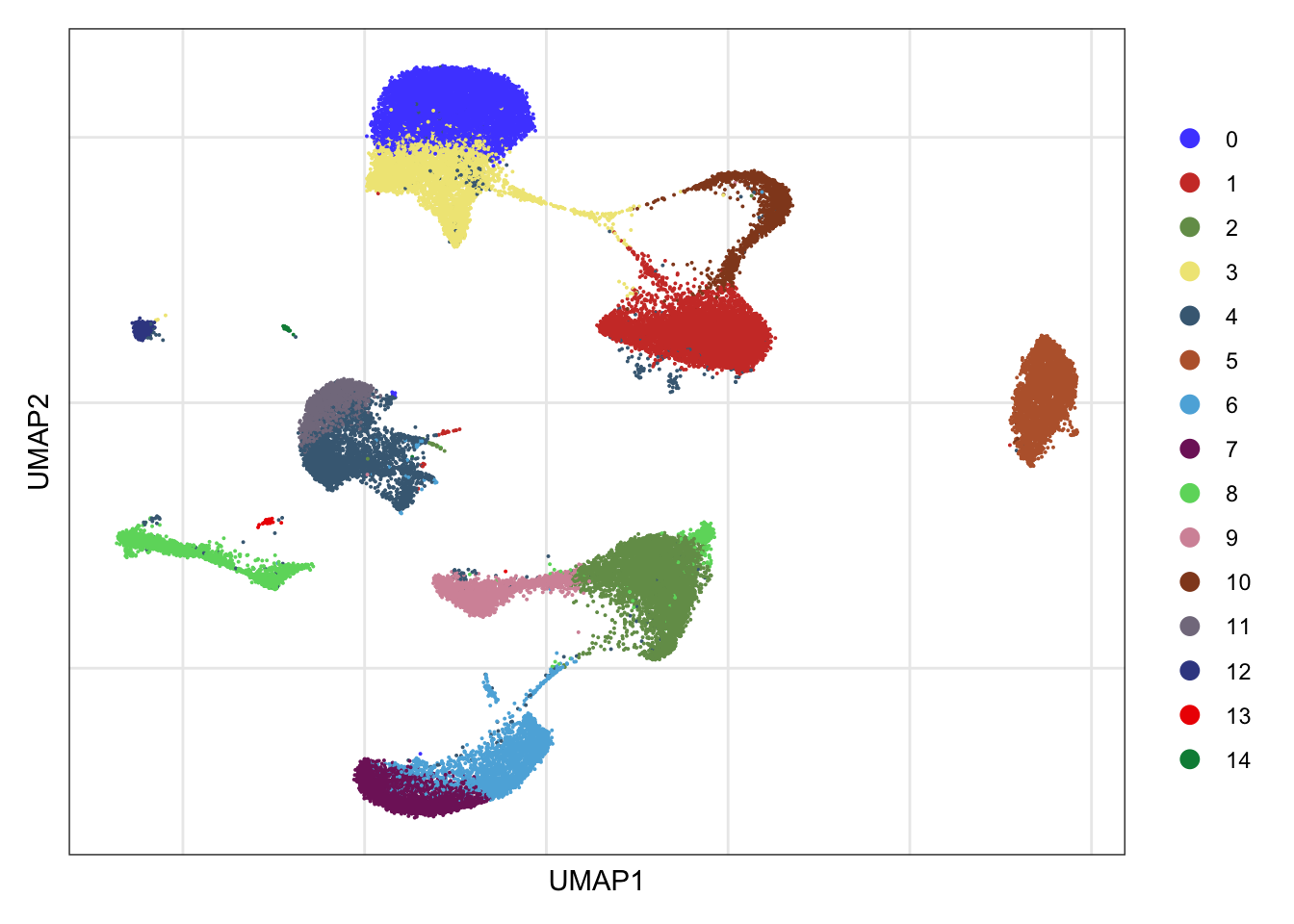
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
DimPlot(seurat, reduction = "umap", cols=colPal,
shuffle = T)+
theme_void()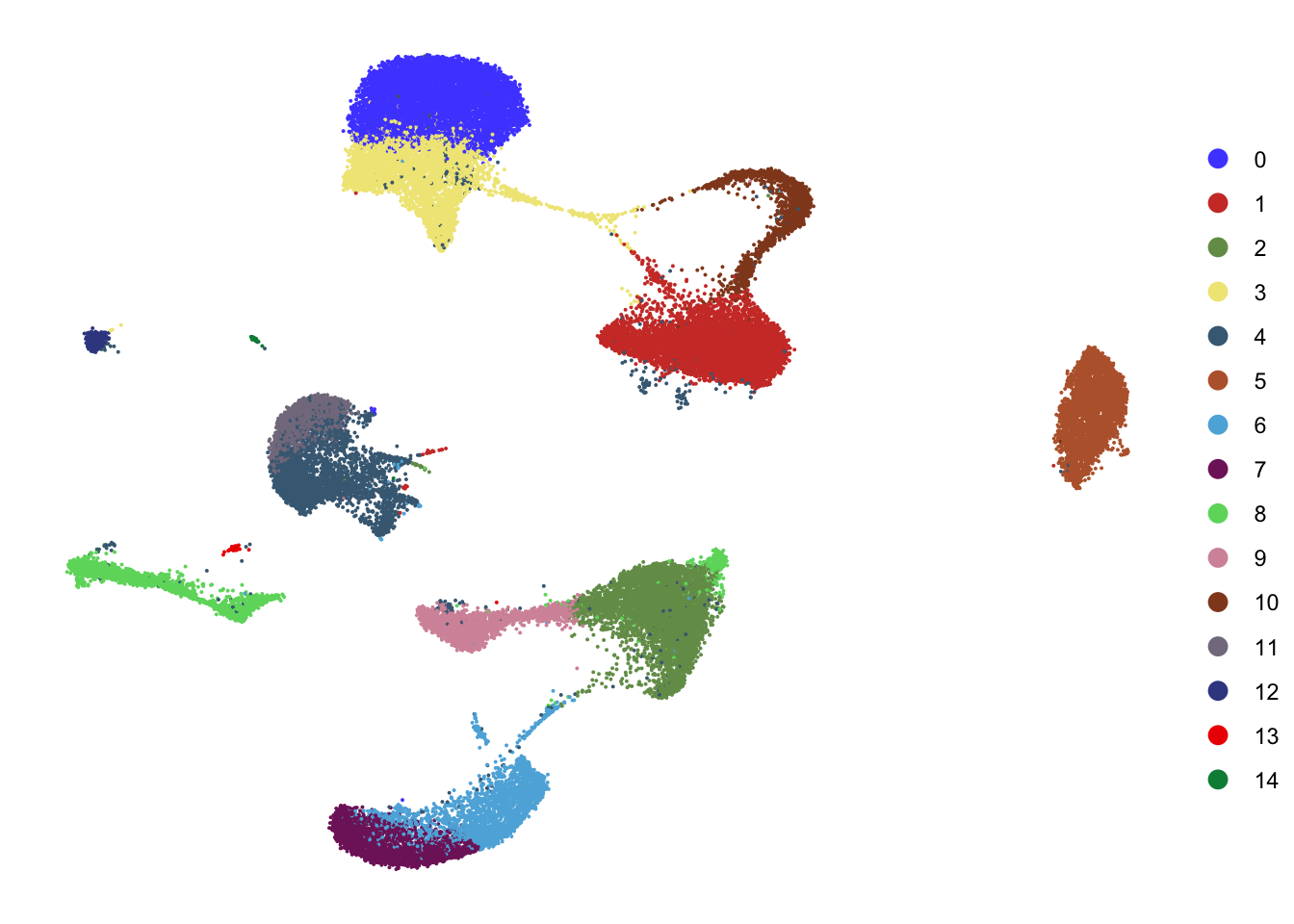
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
label
DimPlot(seurat, reduction = "umap", group.by = "label", cols=colLab)+
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
panel.grid.minor = element_blank()) +
xlab("UMAP1") +
ylab("UMAP2")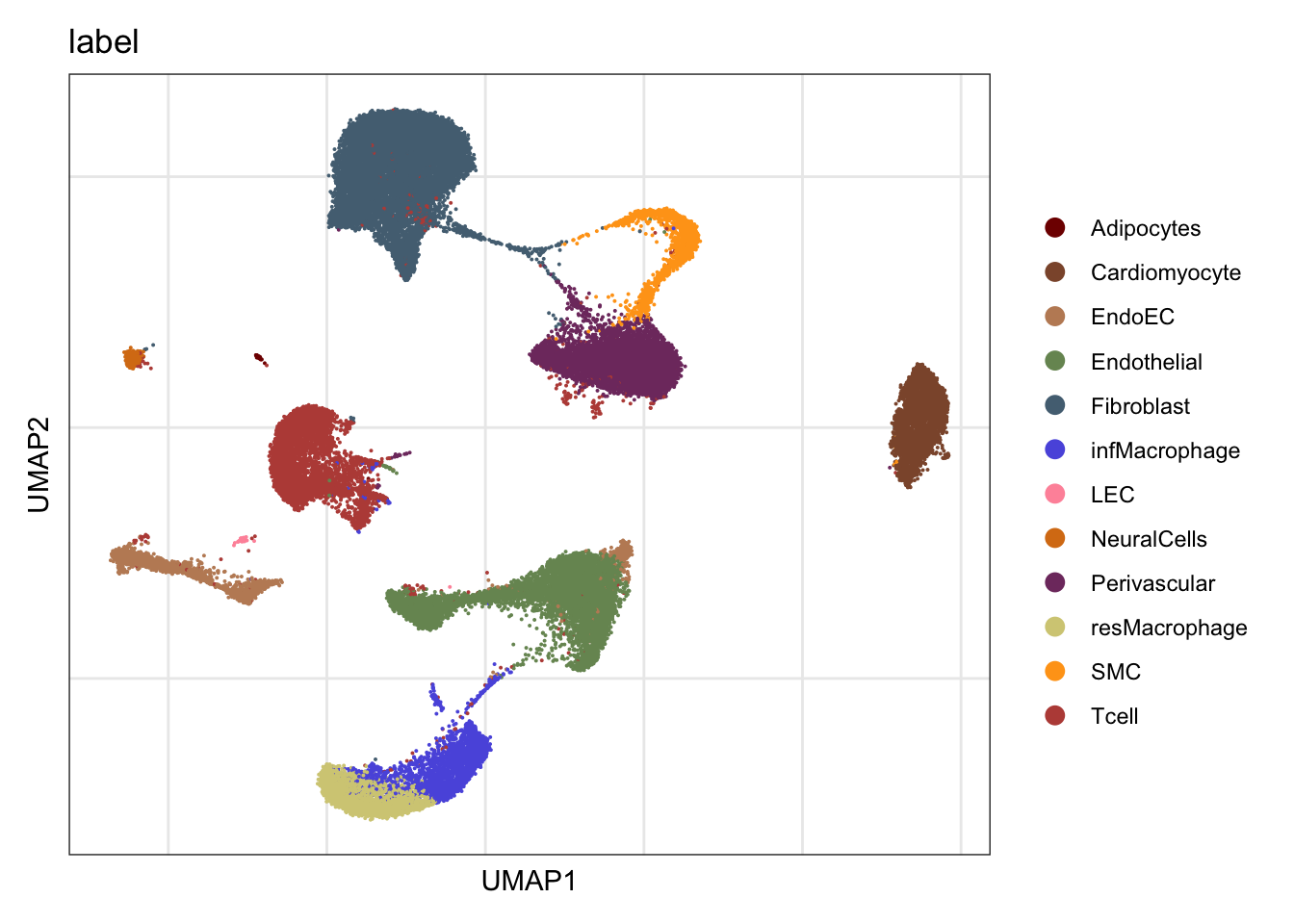
DimPlot(seurat, reduction = "umap", group.by = "label", cols=colLab,
shuffle = T)+
theme_void()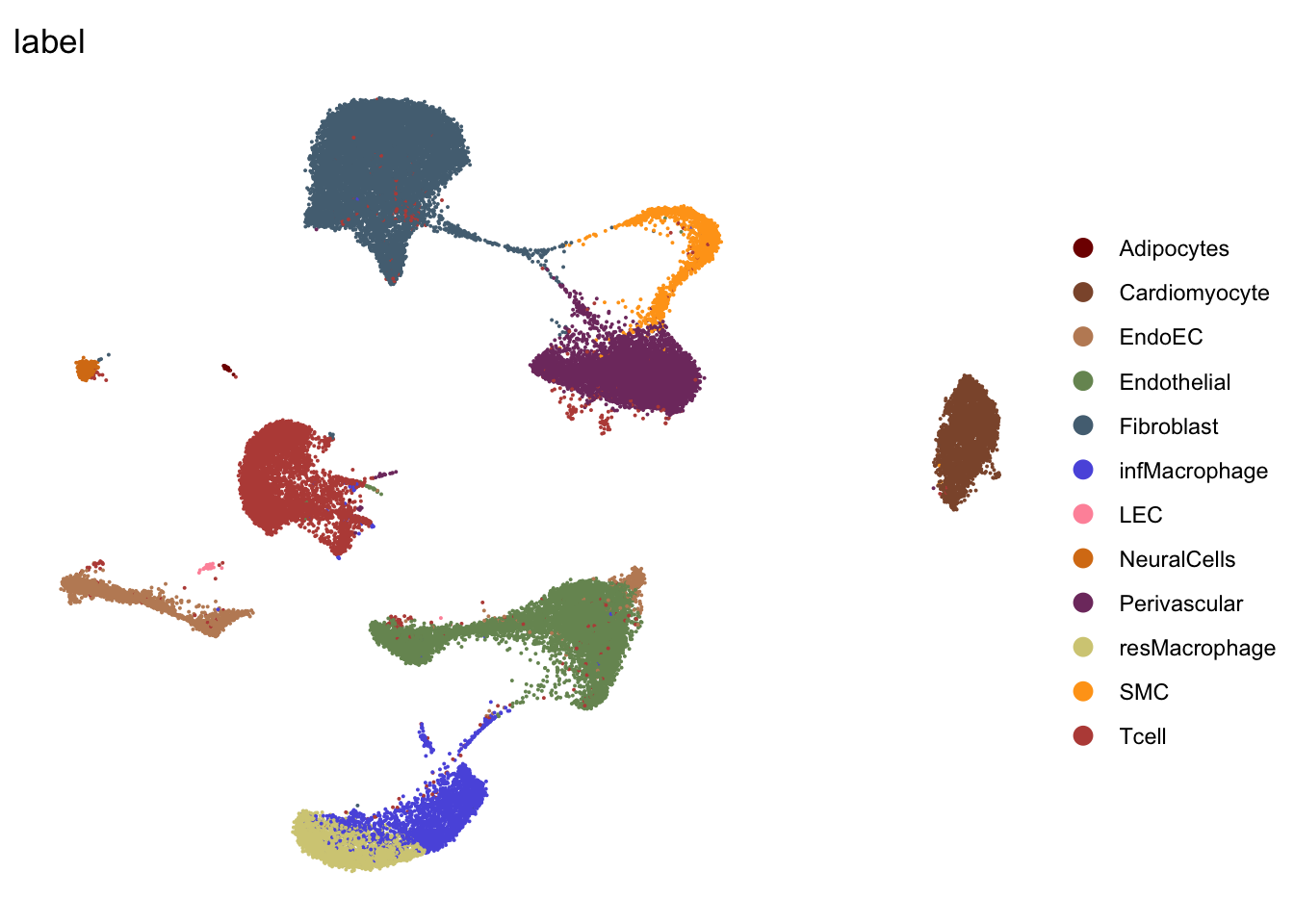
ID
DimPlot(seurat, reduction = "umap", group.by = "ID", cols=colID, shuffle = T)+
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
panel.grid.minor = element_blank()) +
xlab("UMAP1") +
ylab("UMAP2")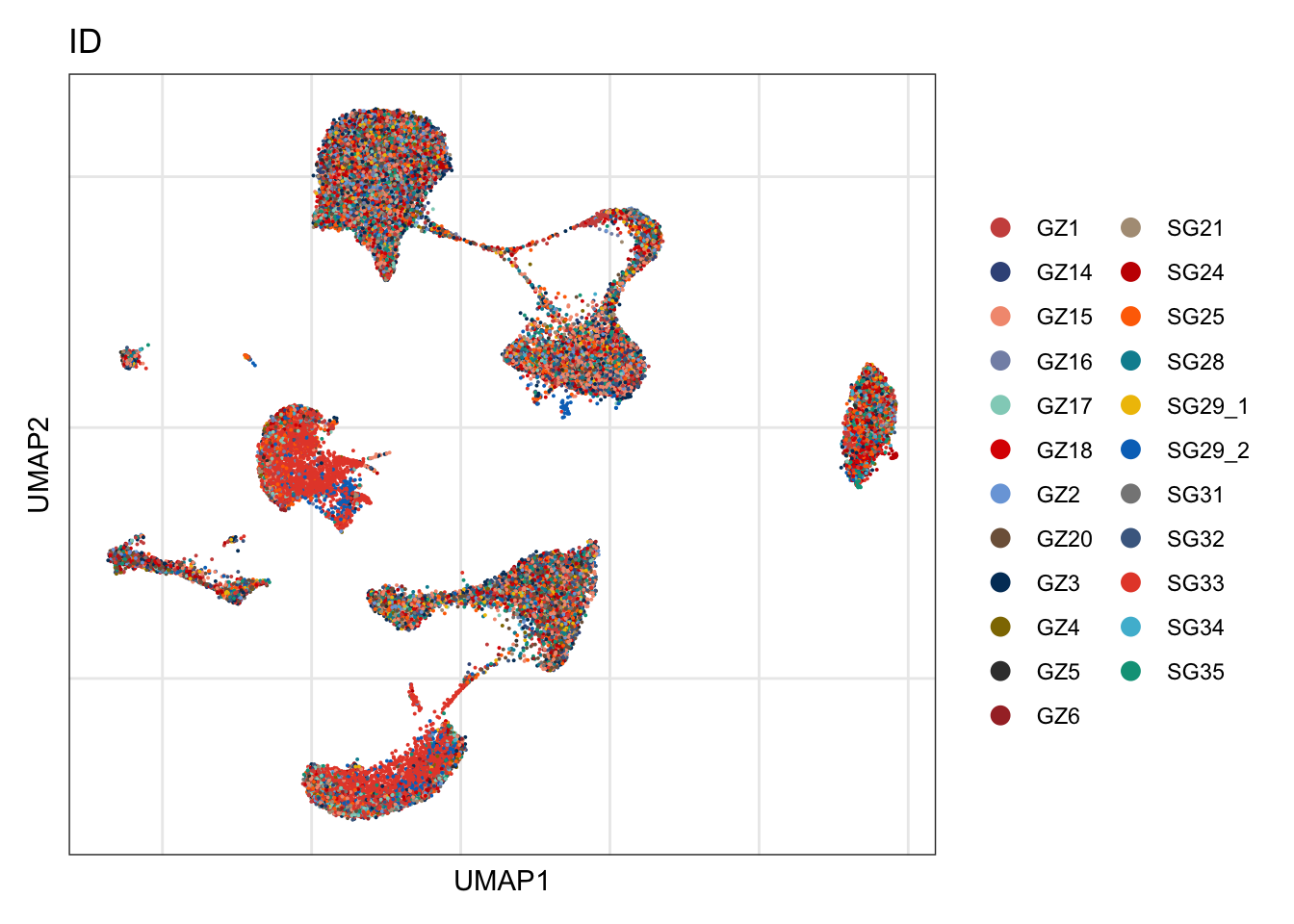
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
DimPlot(seurat, reduction = "umap", group.by = "ID", cols=colID,
shuffle = T)+
theme_void()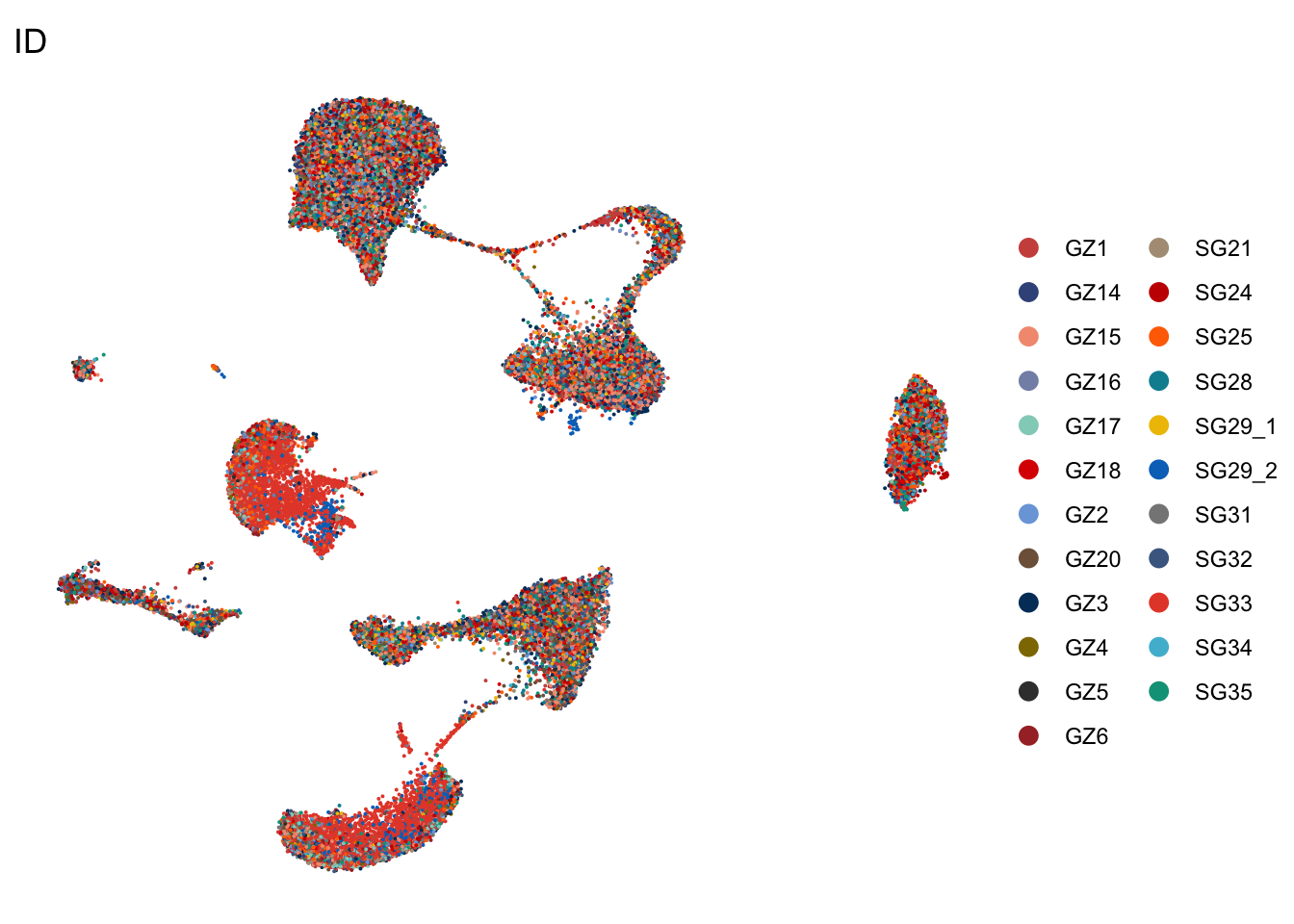
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
cond
DimPlot(seurat, reduction = "umap", group.by = "cond", cols=colCond)+
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
panel.grid.minor = element_blank()) +
xlab("UMAP1") +
ylab("UMAP2")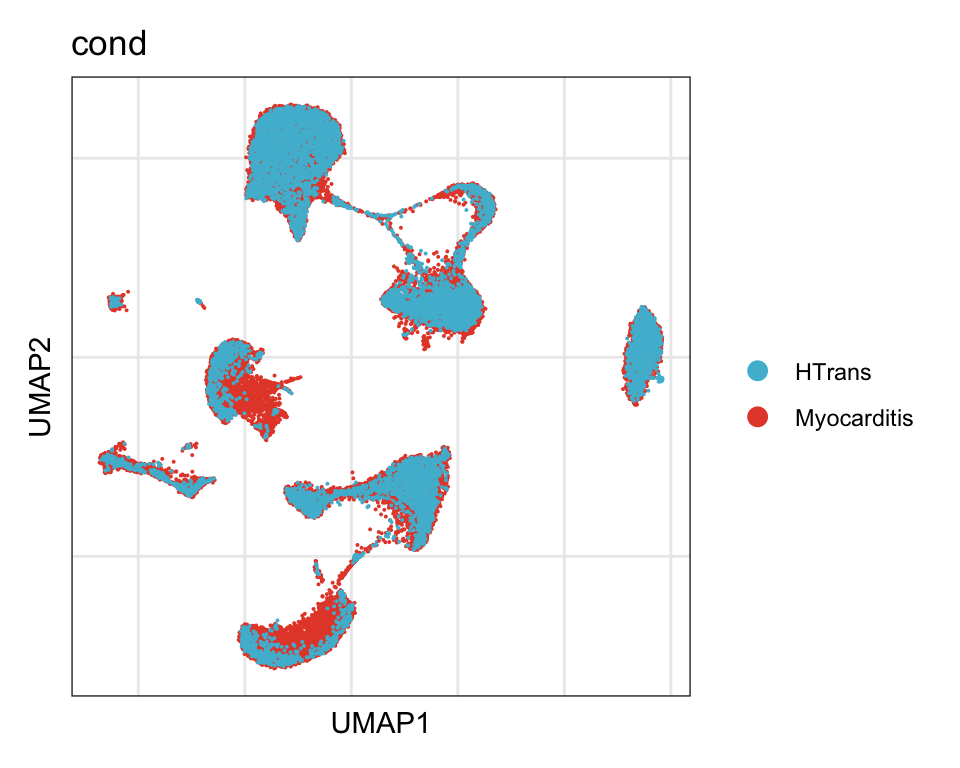
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
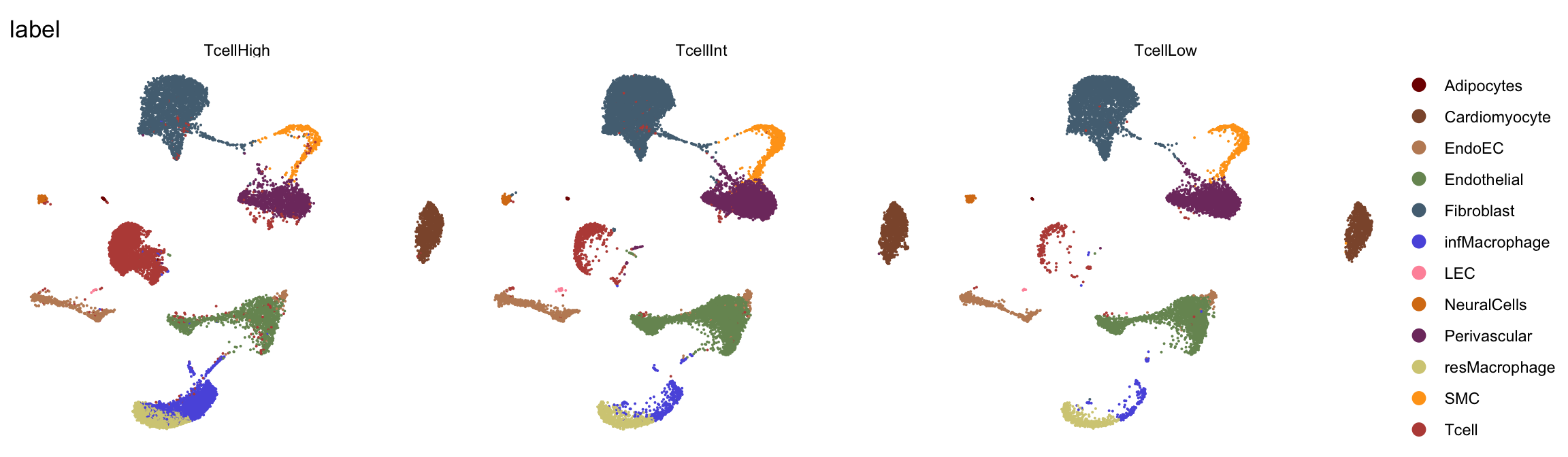
T cell grp
DimPlot(seurat, reduction = "umap", group.by = "TcellGrp", cols=colTgrp)+
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
panel.grid.minor = element_blank()) +
xlab("UMAP1") +
ylab("UMAP2")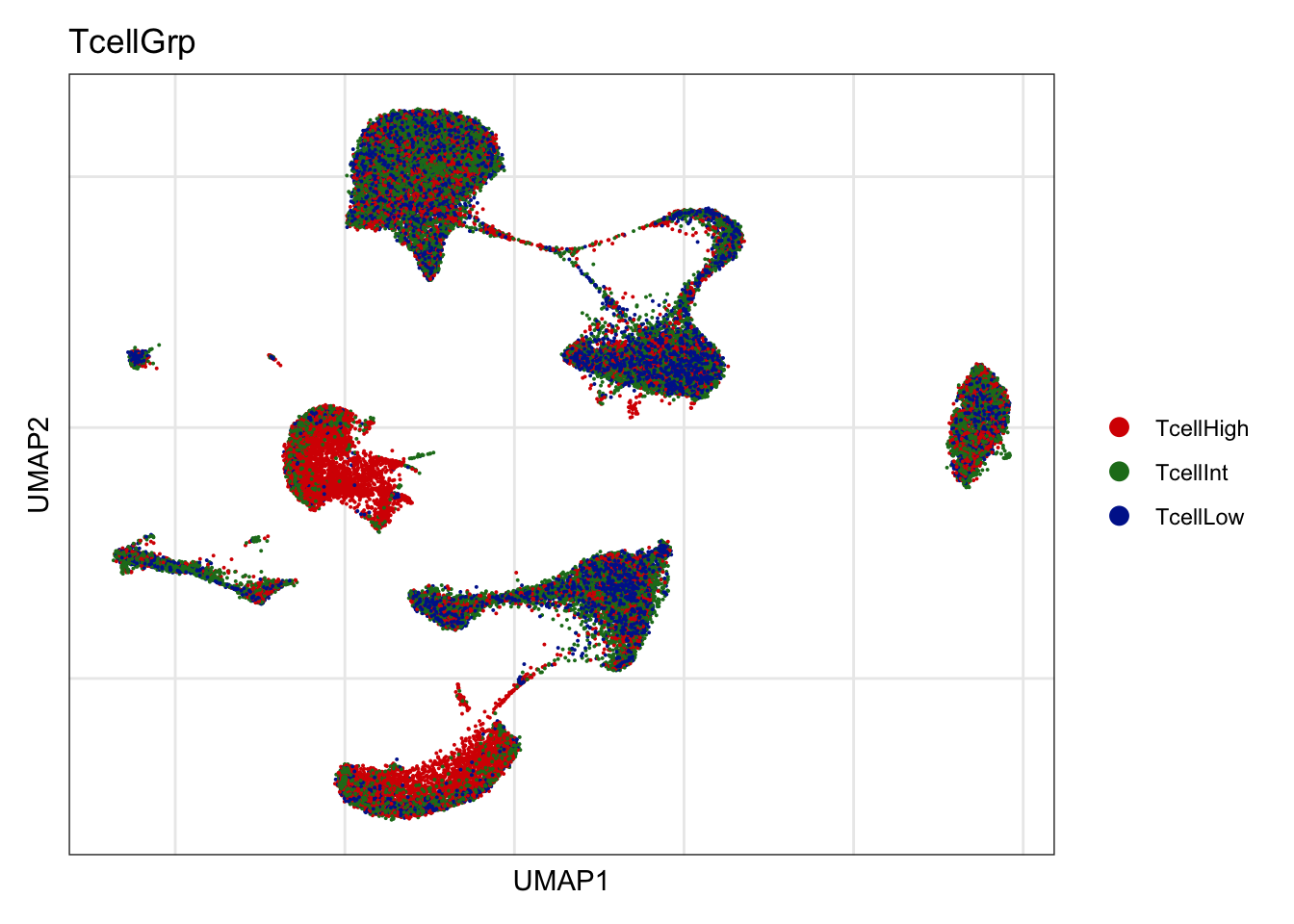
DimPlot(seurat, reduction = "umap", group.by = "TcellGrp", cols=colTgrp,
shuffle = T)+
theme_void()
project signatures
signature cut 2
signDat <- read_delim(file = paste0(basedir,
"/data/GSEA/selGenesSignature.txt"),
delim = "\t")
genes <- data.frame(geneID=rownames(seurat)) %>%
mutate(gene=gsub("^.*\\.", "", geneID))
signDat <- signDat %>% left_join(.,genes, by="gene")
allSign <- unique(signDat$signature)
sce <- as.SingleCellExperiment(seurat)
## add reduced dim
seurat2 <- seurat
DefaultAssay(object = seurat2) <- "integrated"
sce2 <- as.SingleCellExperiment(seurat2)
reducedDims(sce) <- reducedDims(sce2)
remove(seurat2)
remove(sce2)
treatGrps <- unique(sce$TcellGrp)
cutOff <- 2
pal = viridis(100)
sc <- scale_colour_gradientn(colours = pal, limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]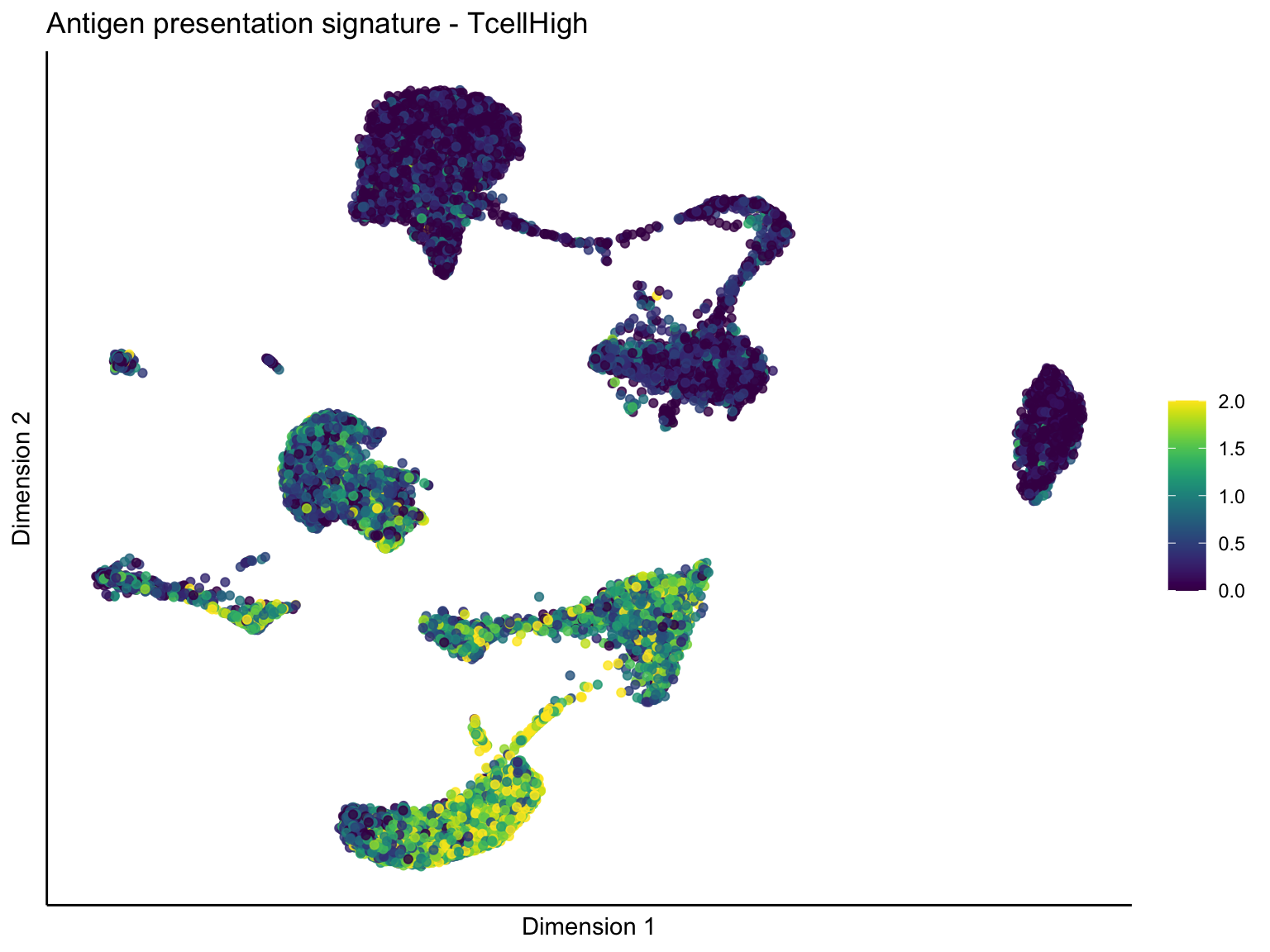
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]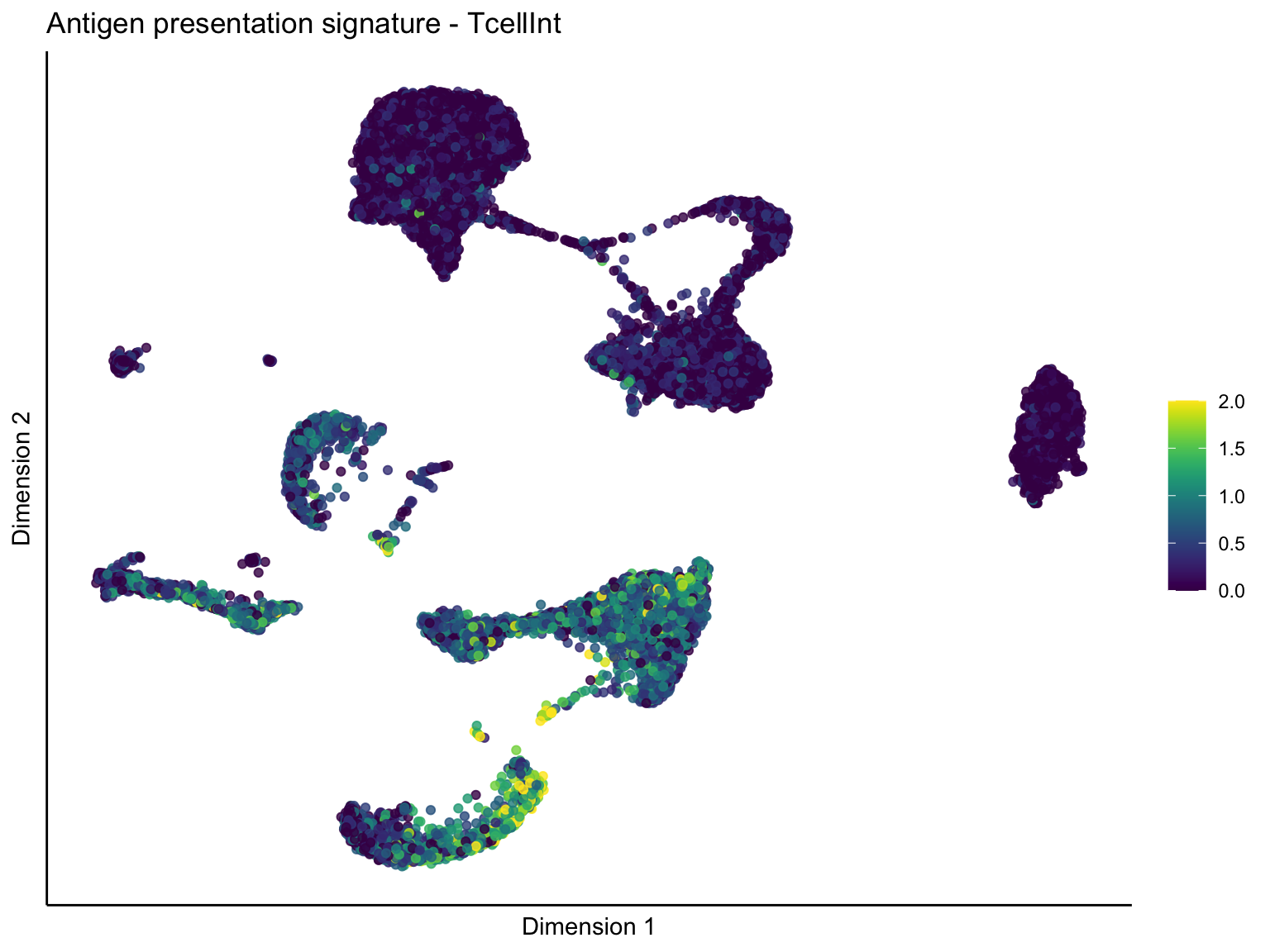
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]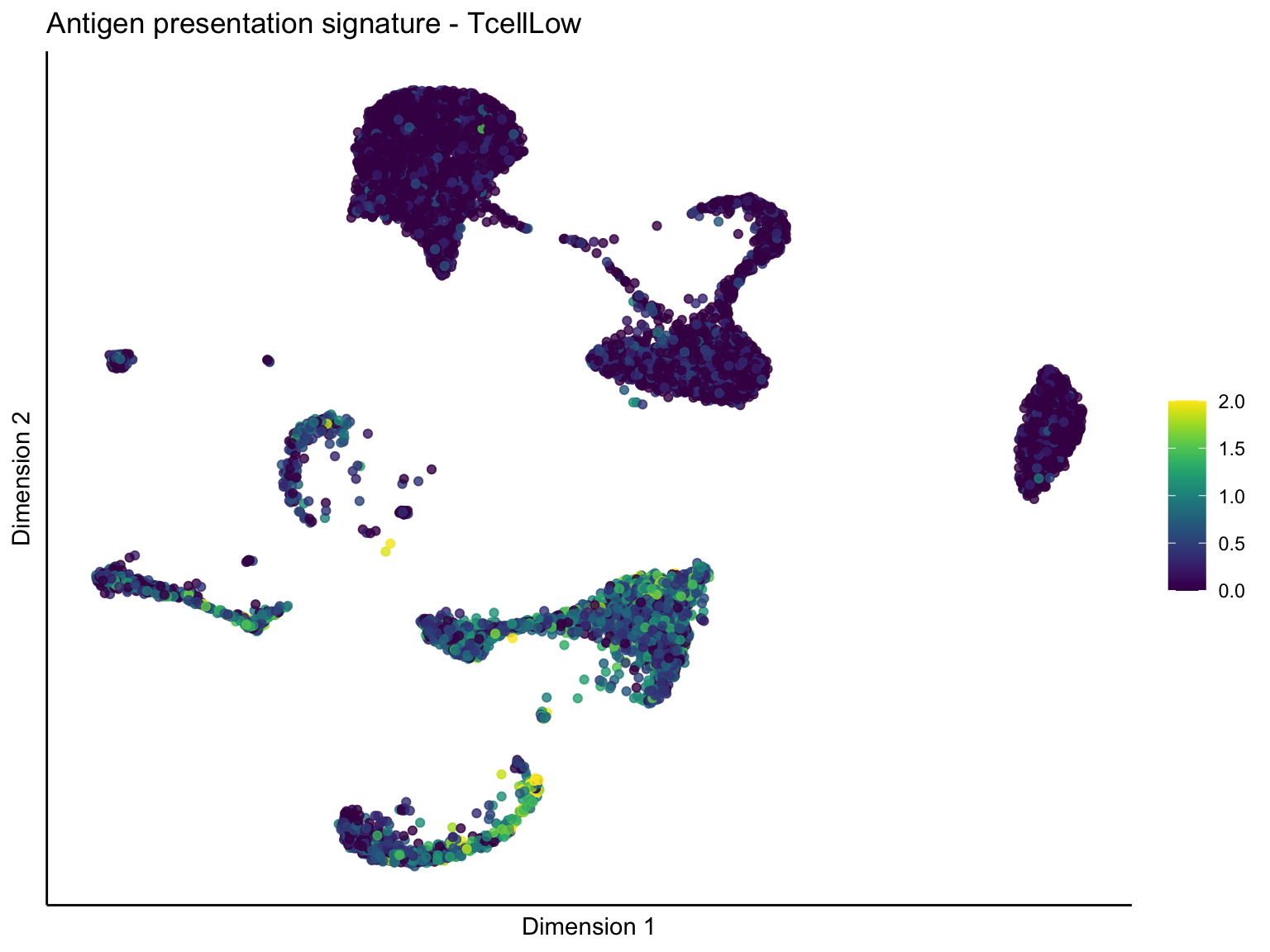
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]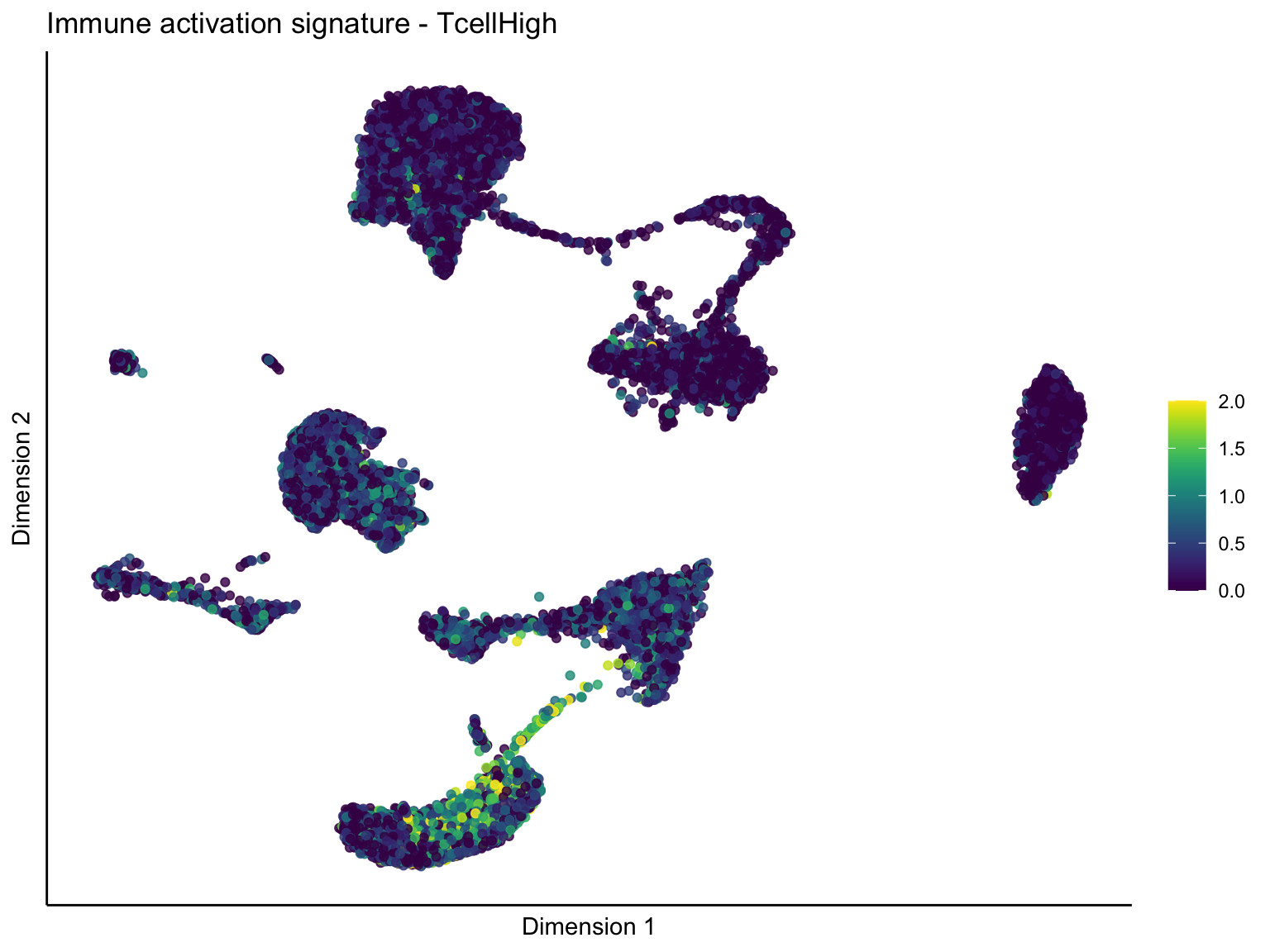
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]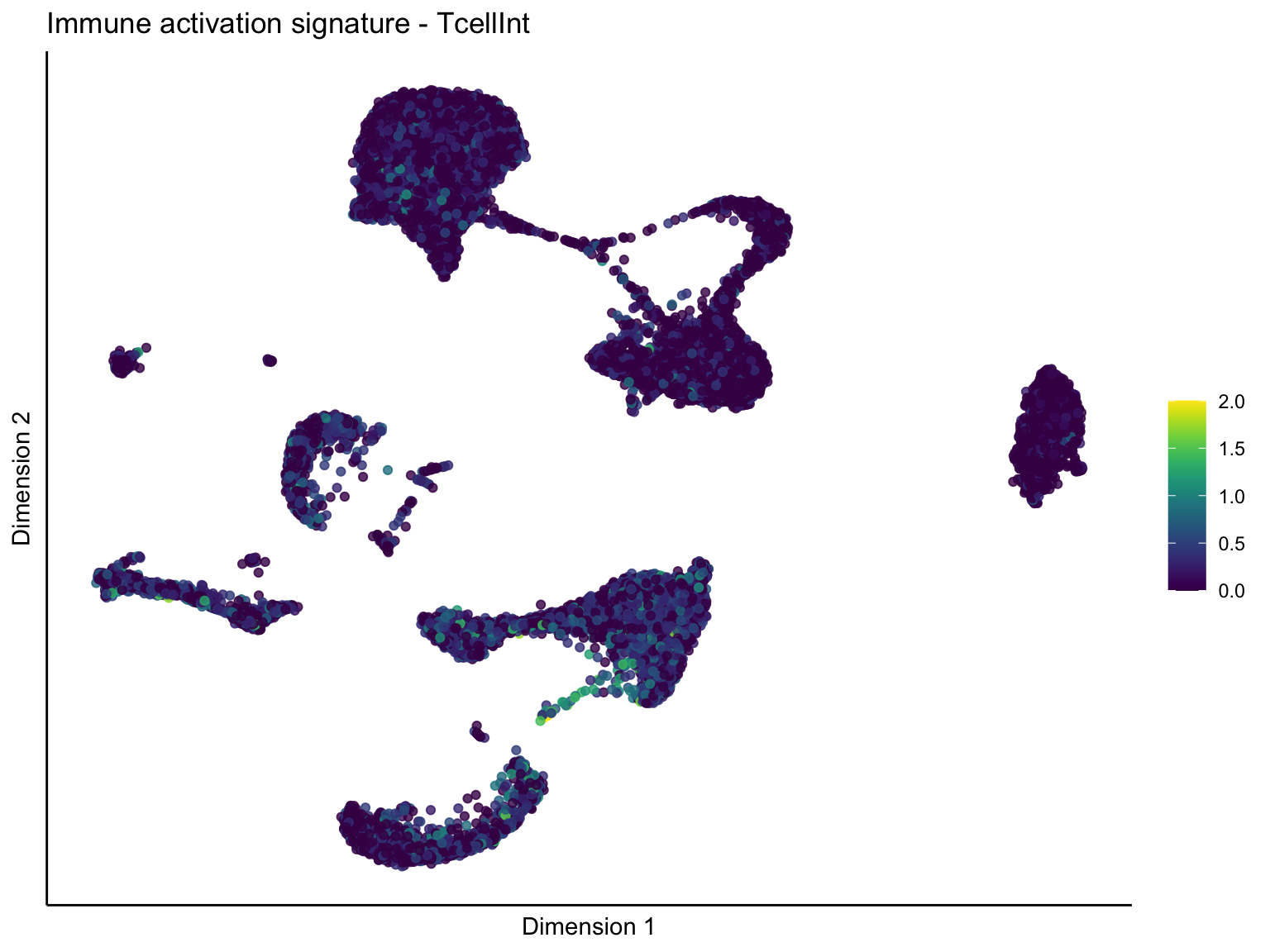
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]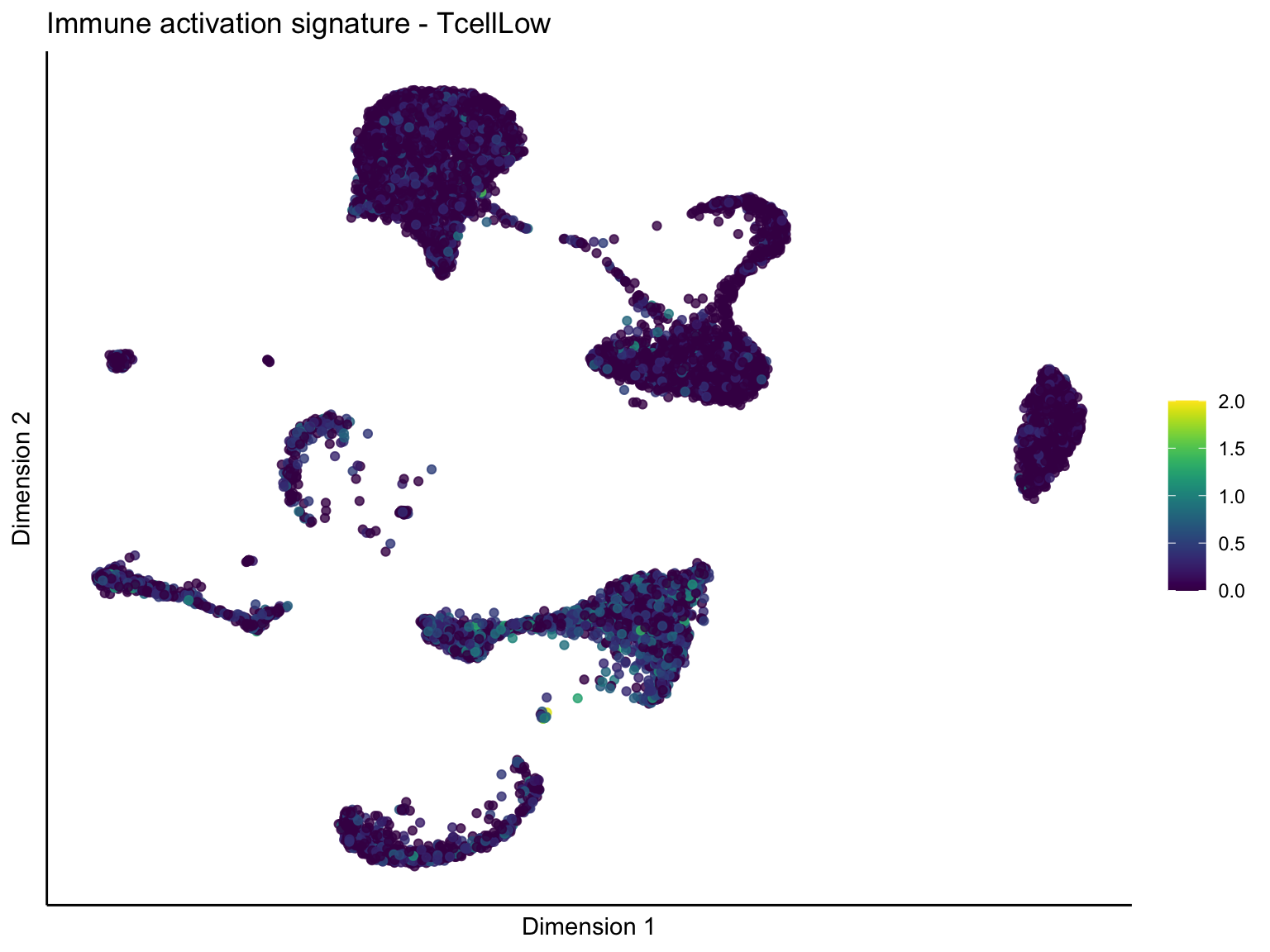
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]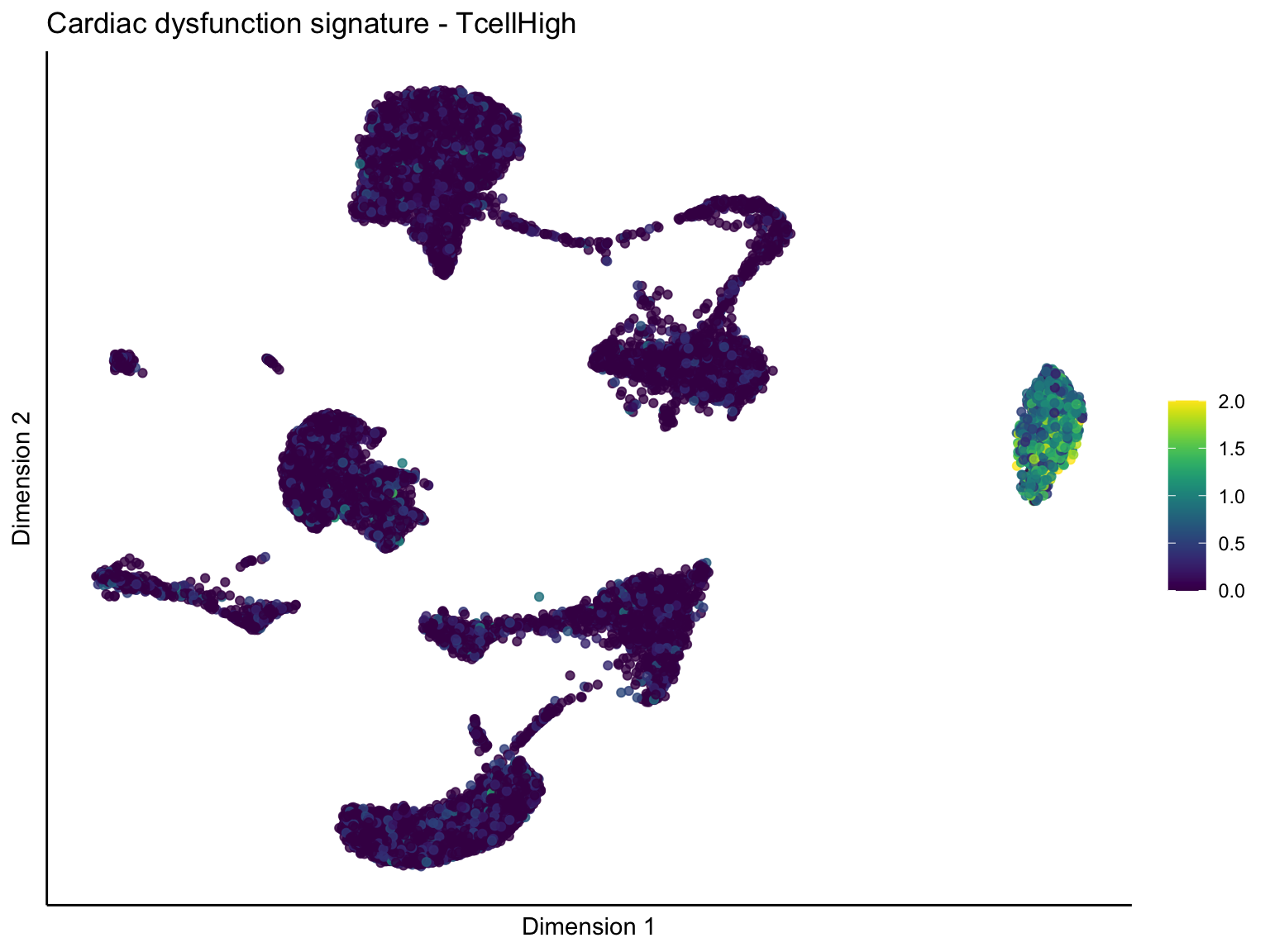
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]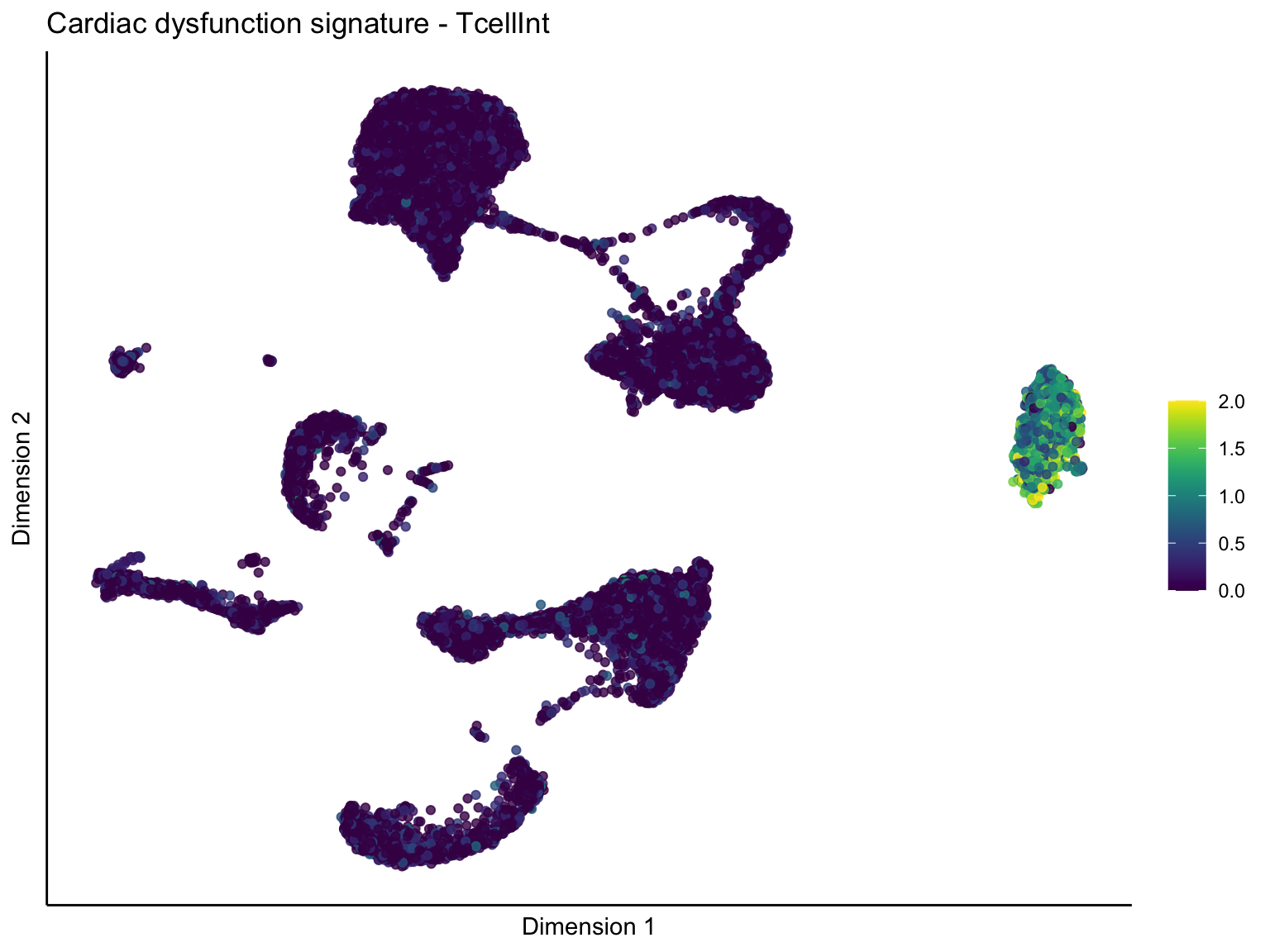
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]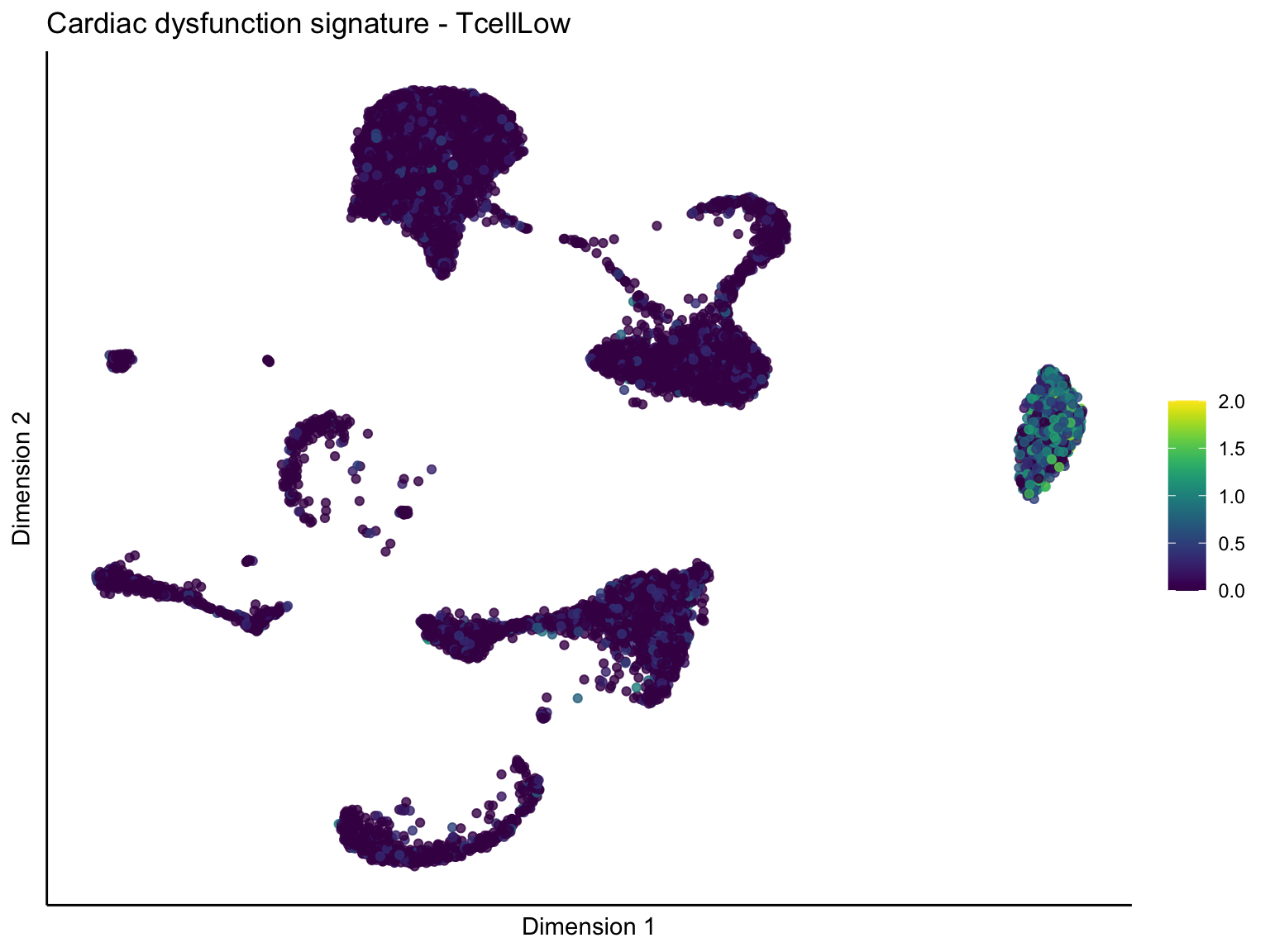
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]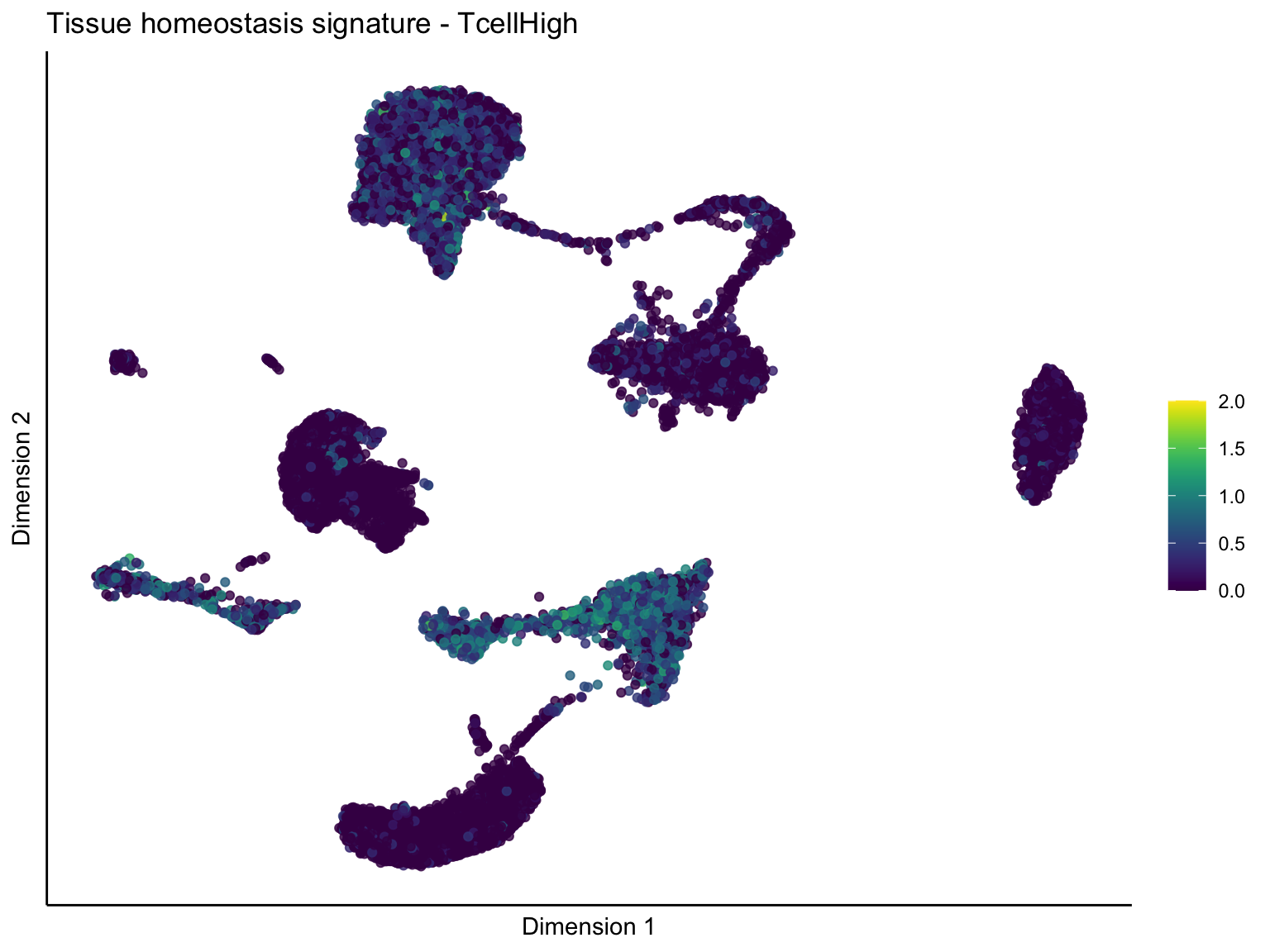
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]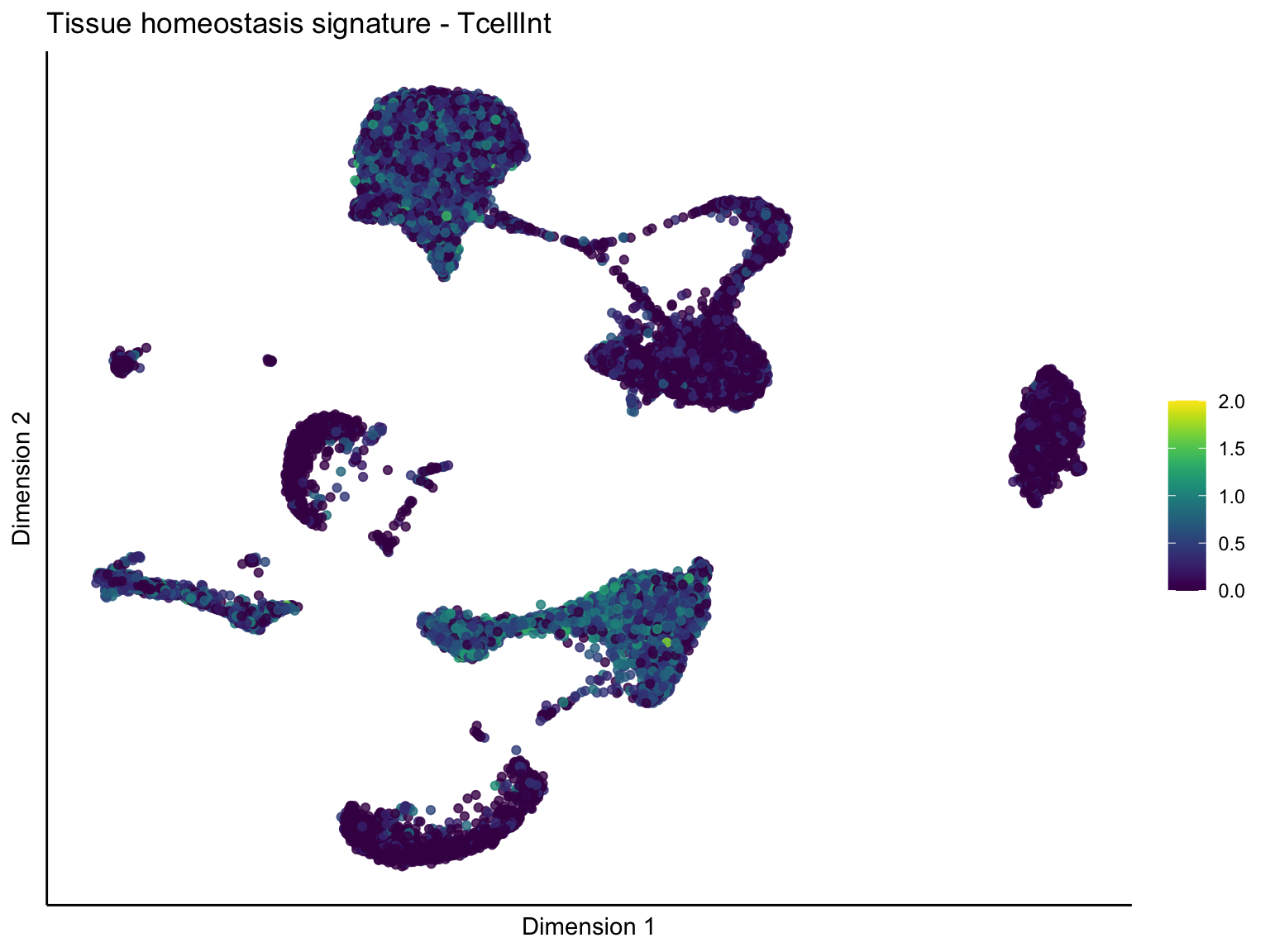
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]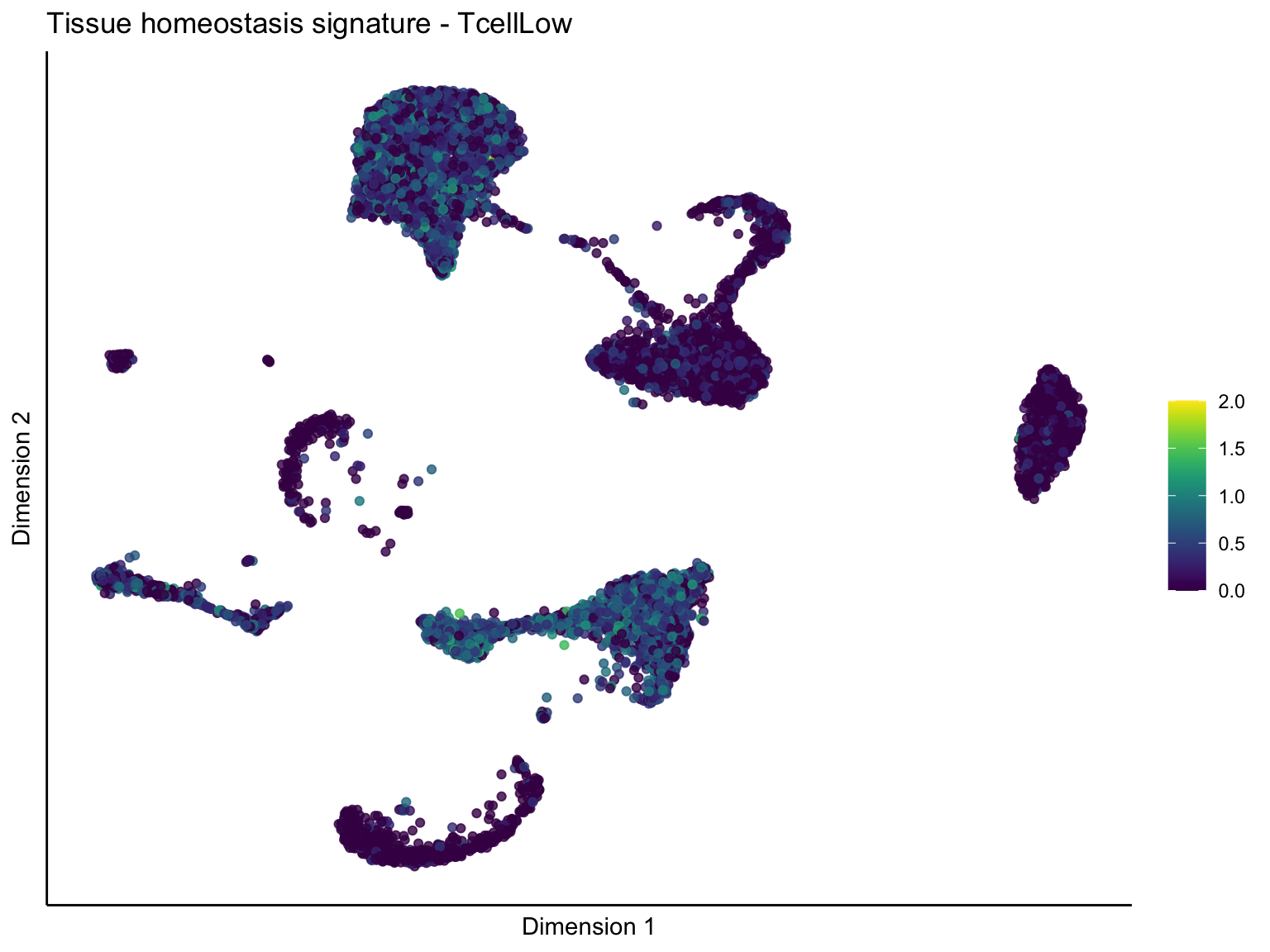
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
pal = colorRampPalette(rev(brewer.pal(11, 'RdBu')))
sc <- scale_colour_gradientn(colours = pal(100), limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]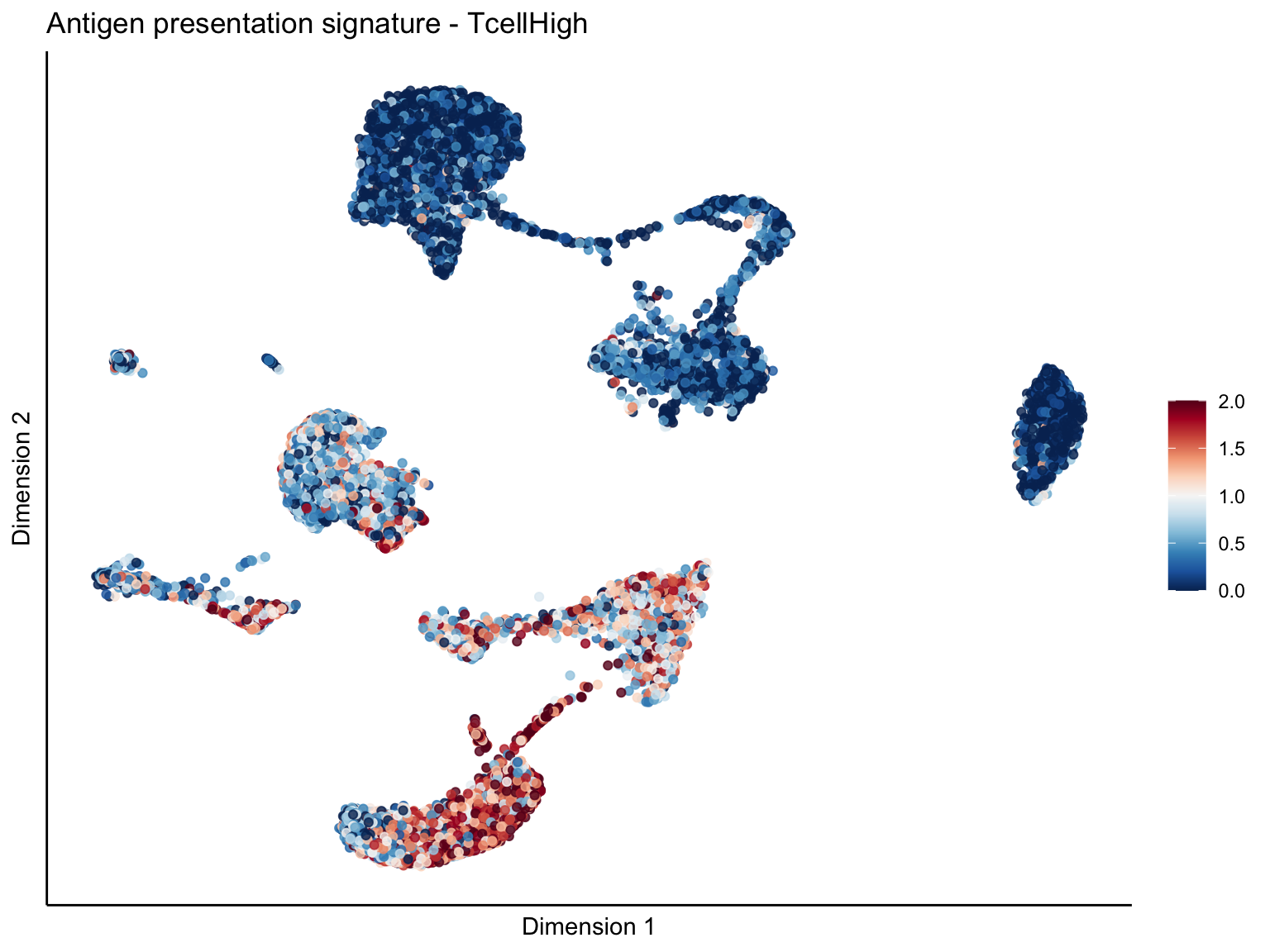
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]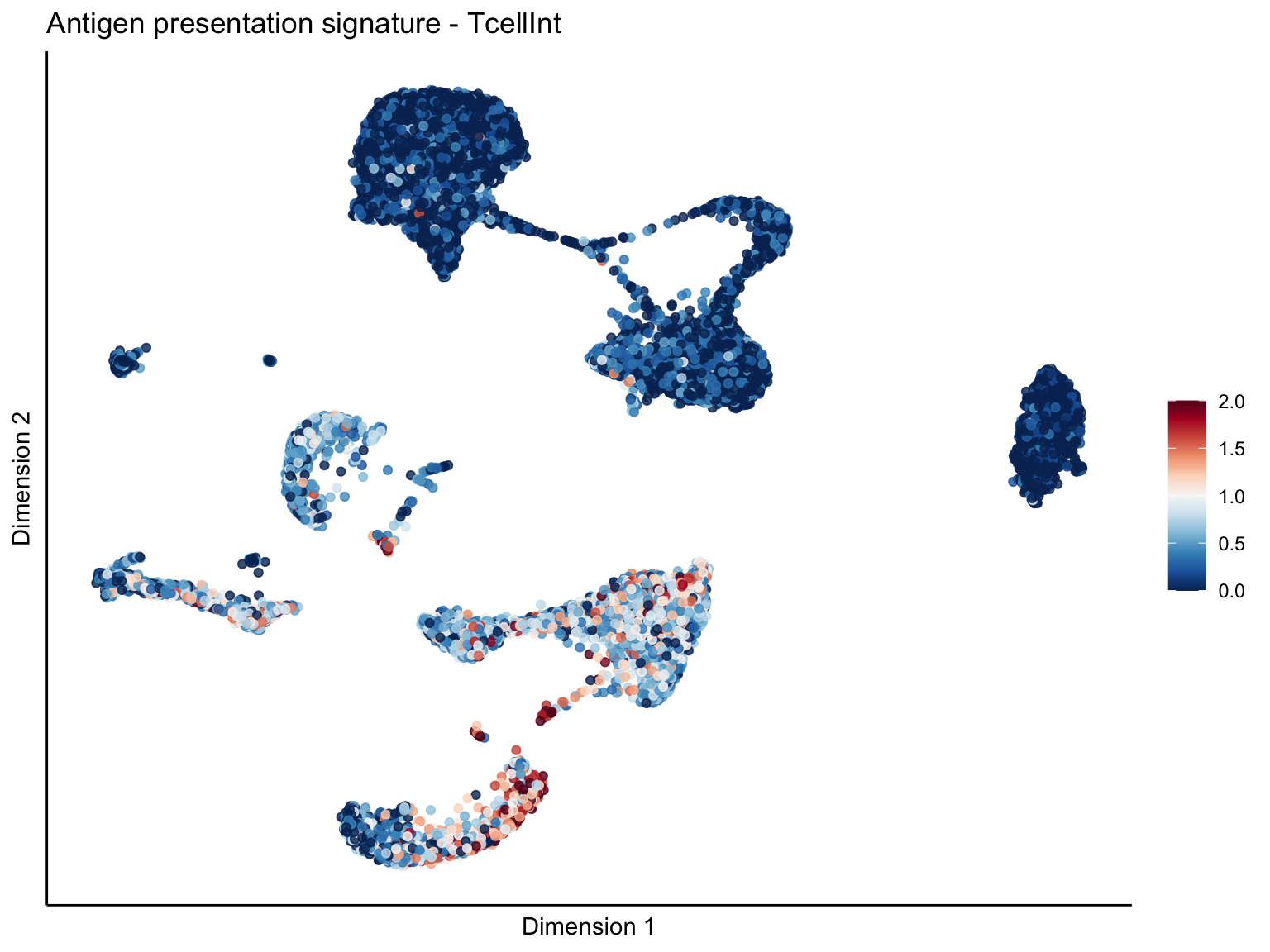
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]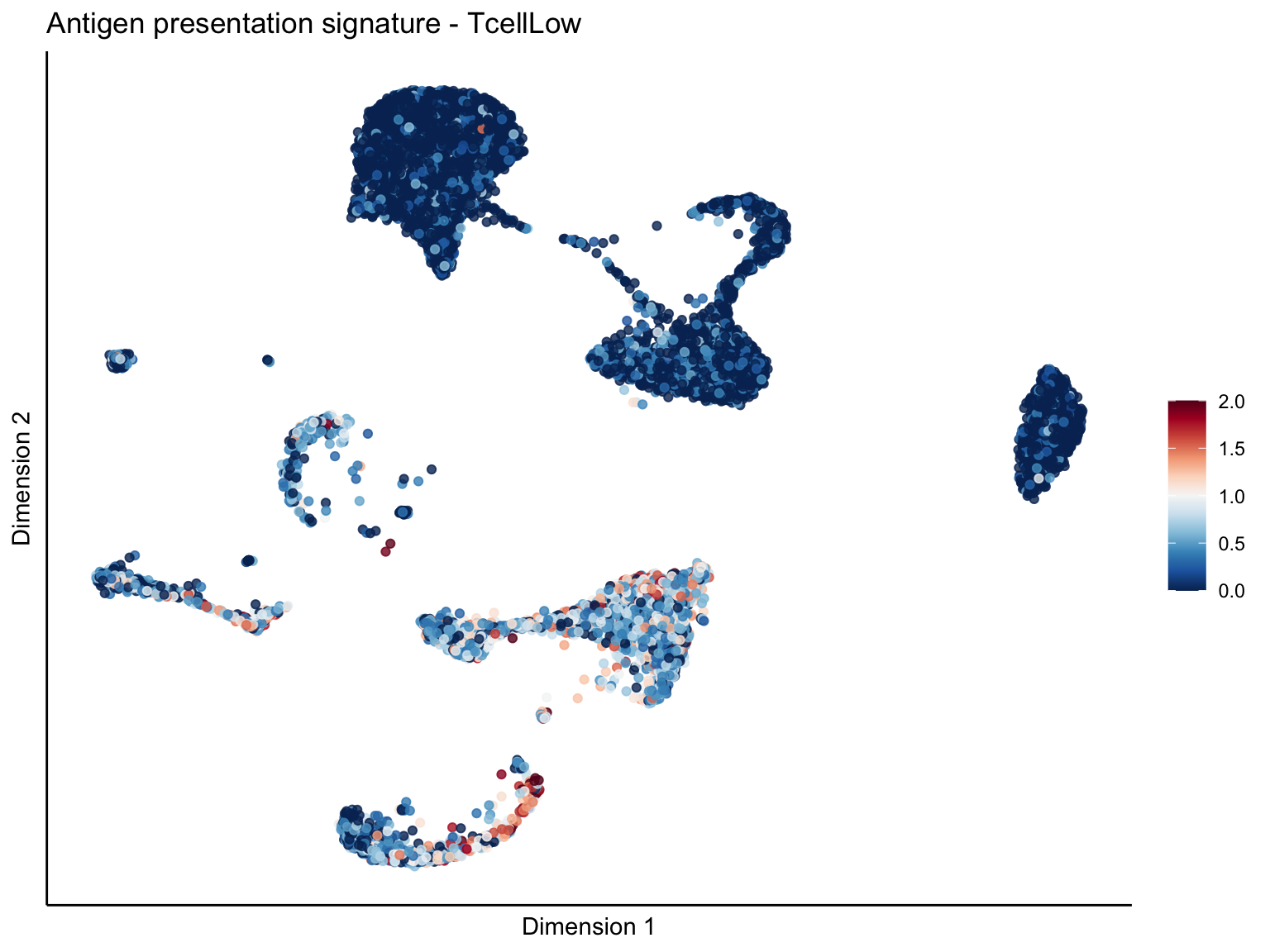
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]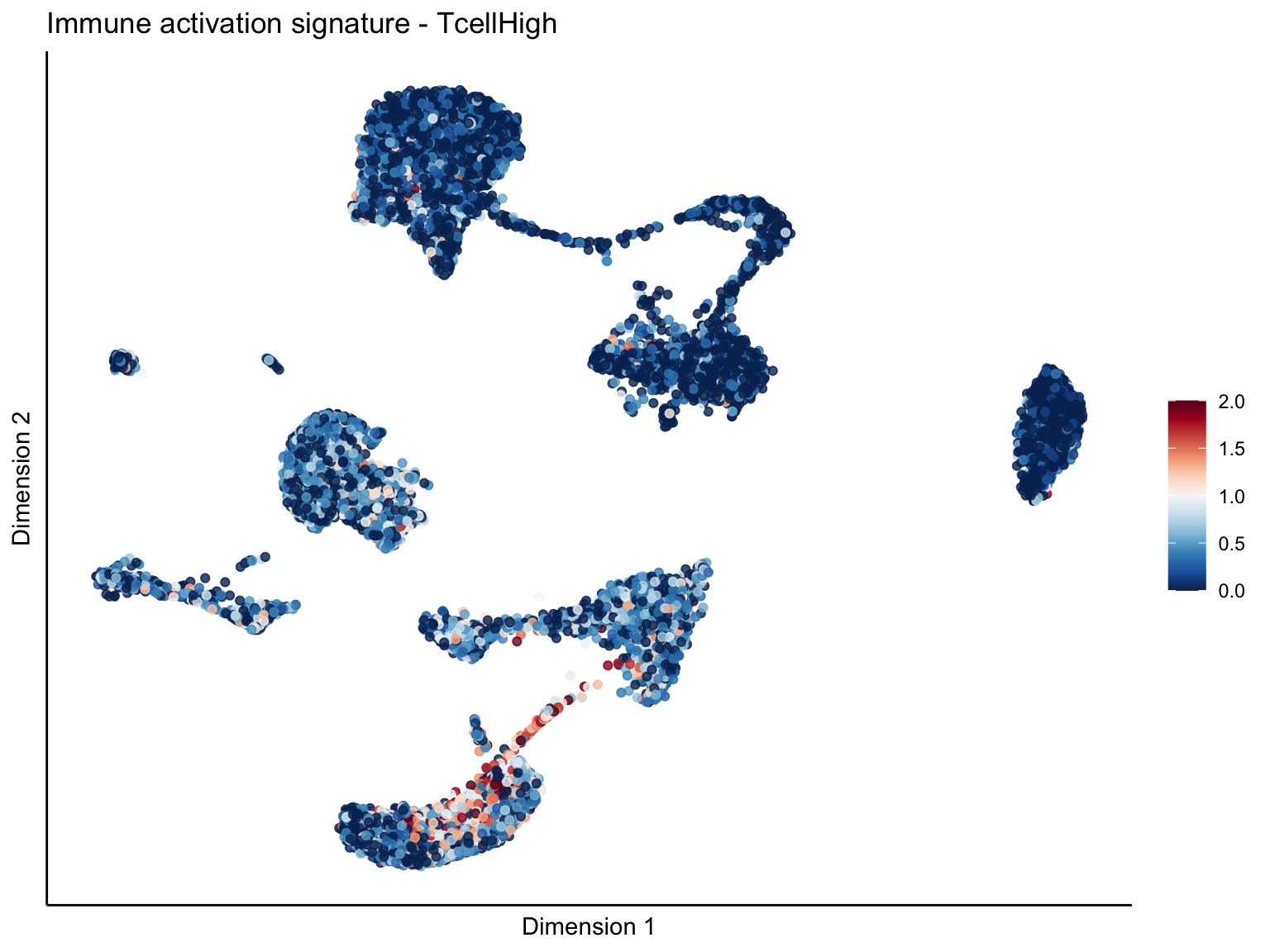
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]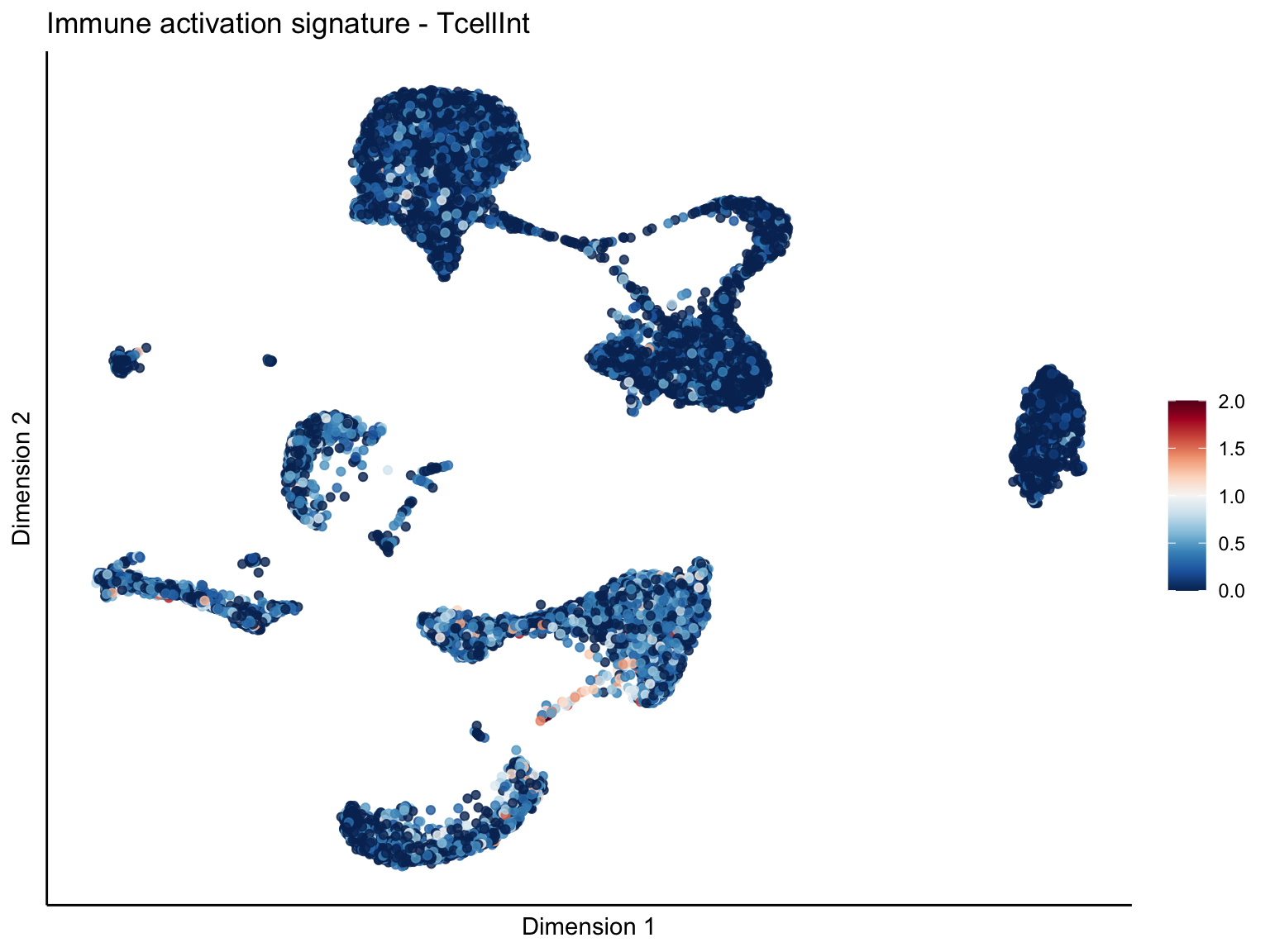
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]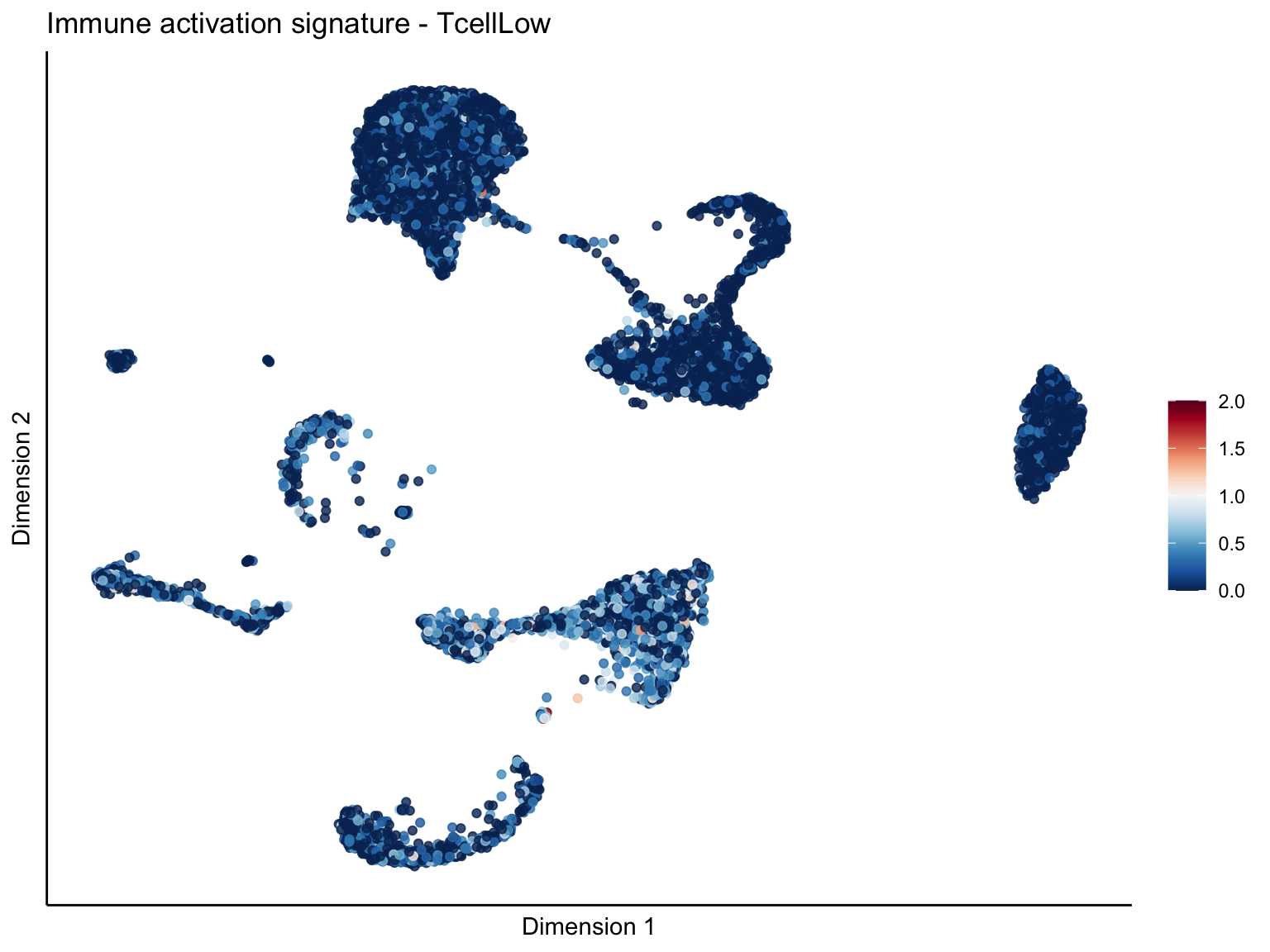
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]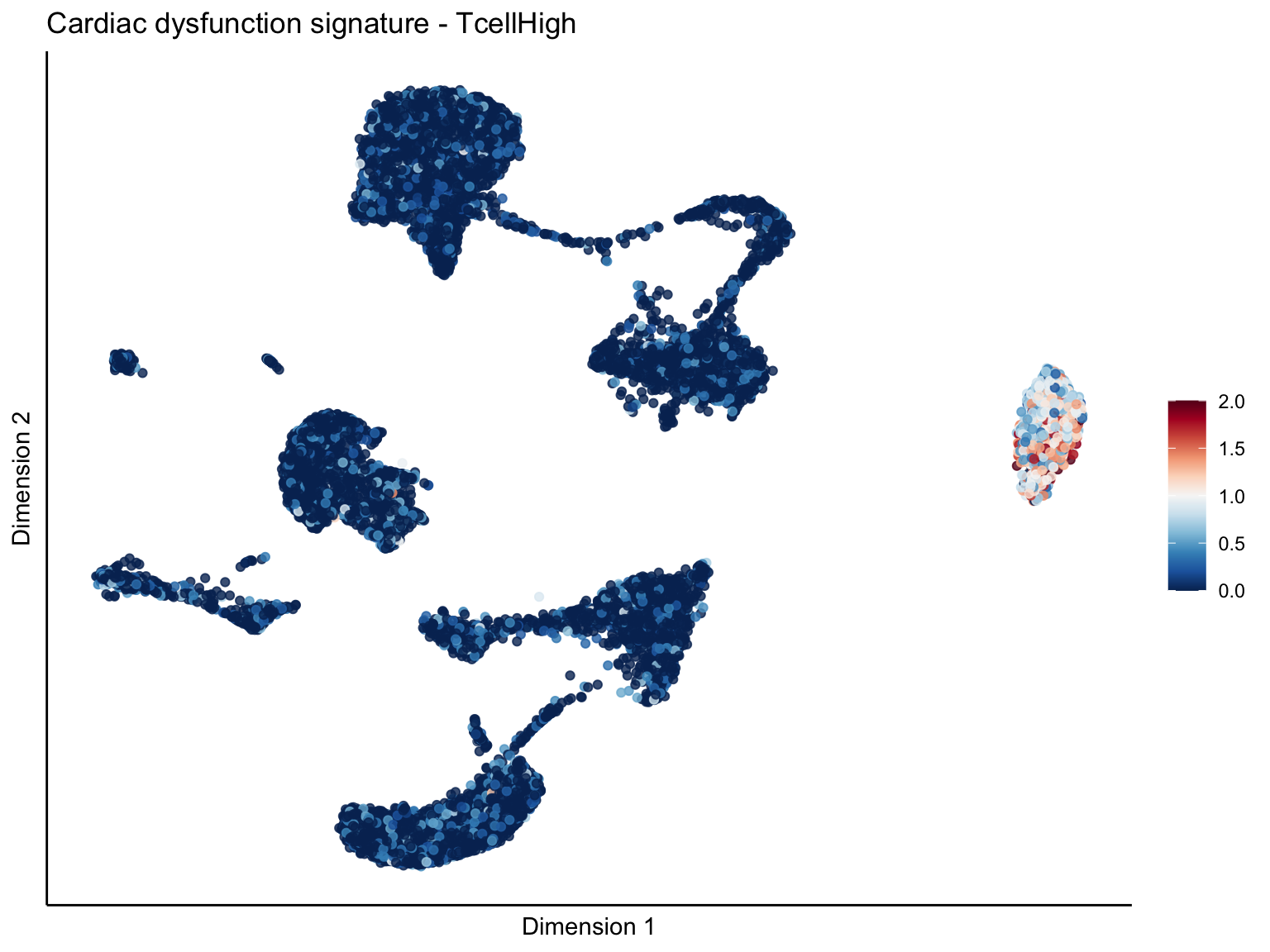
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]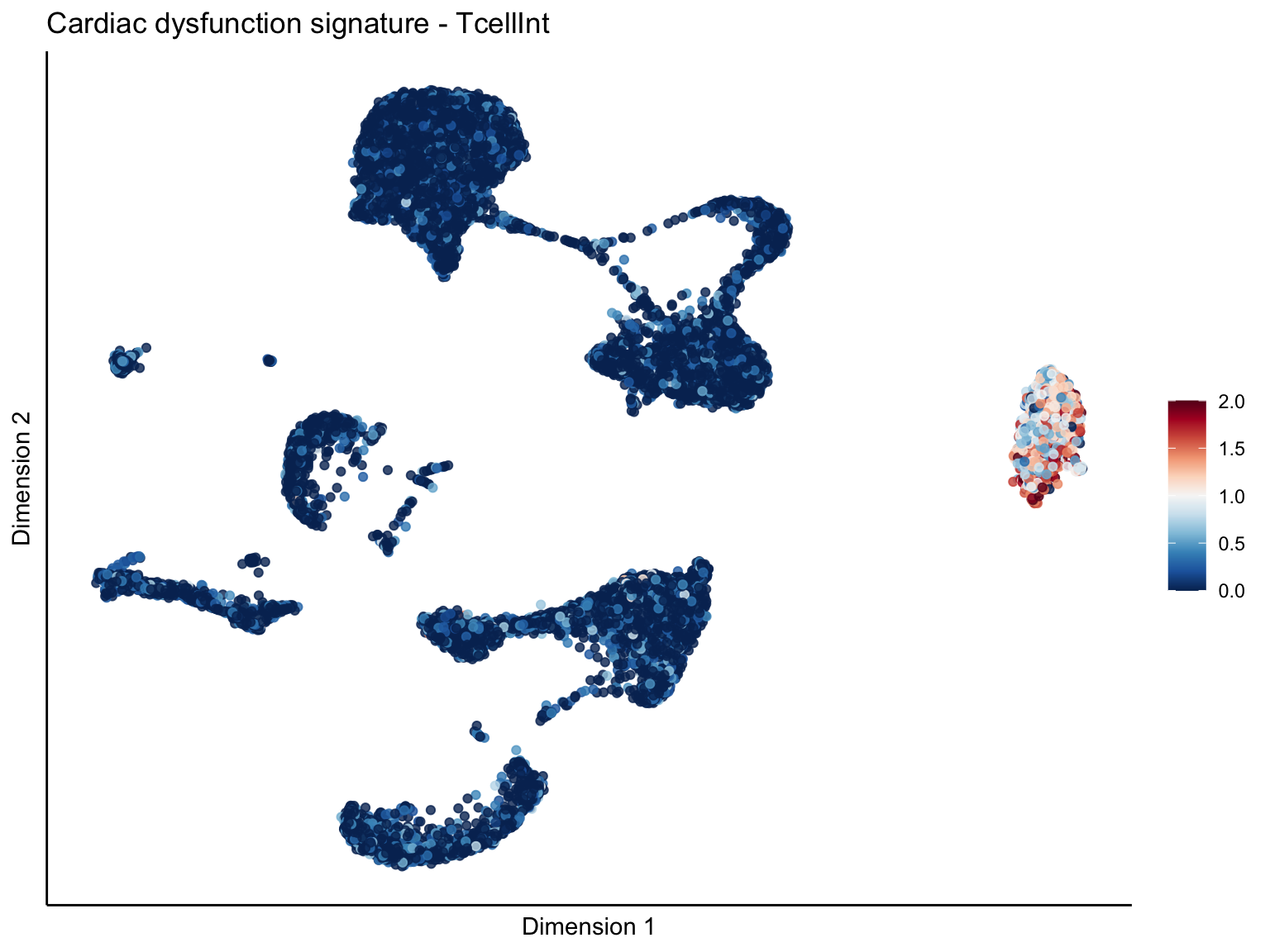
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]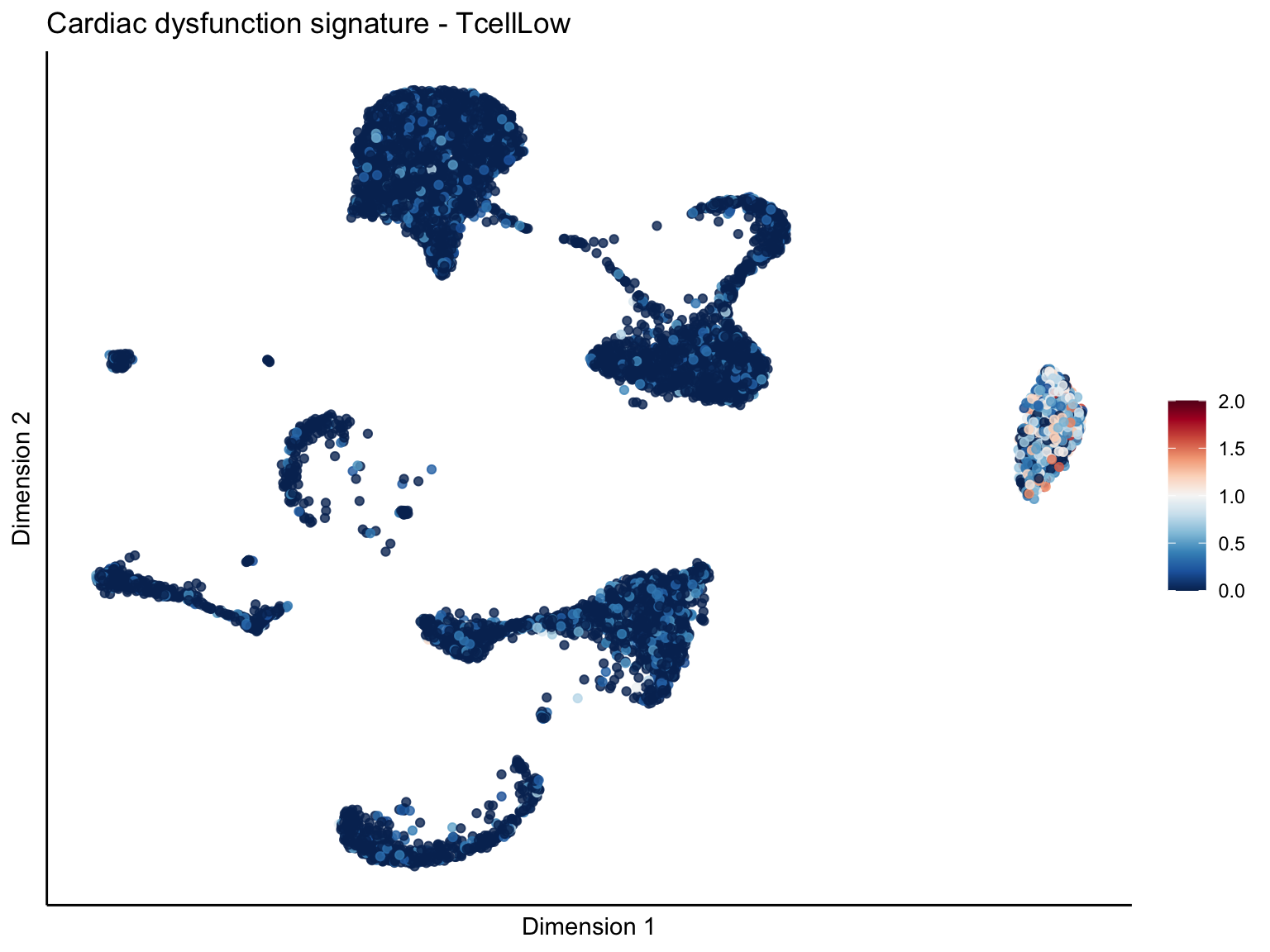
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]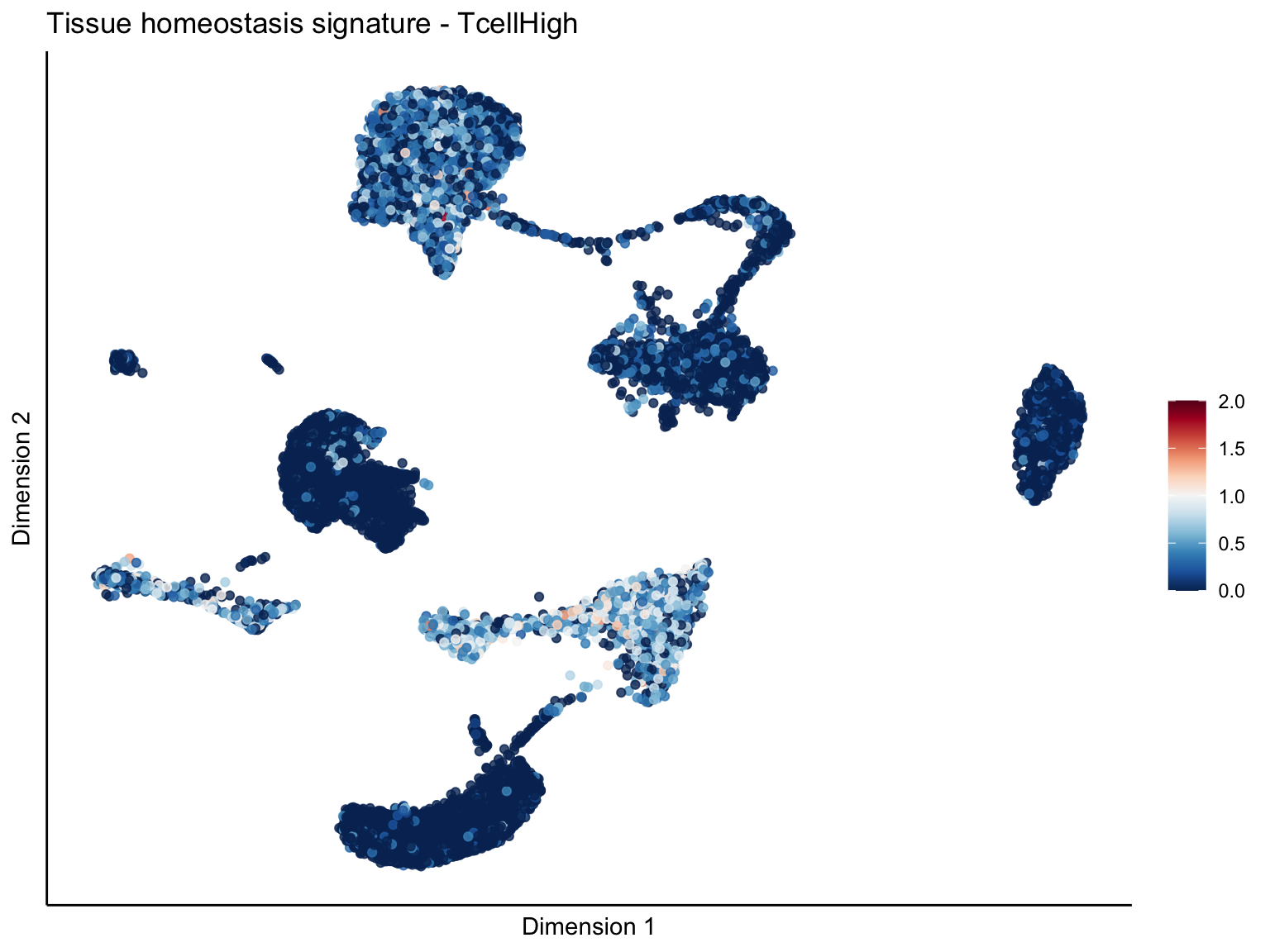
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]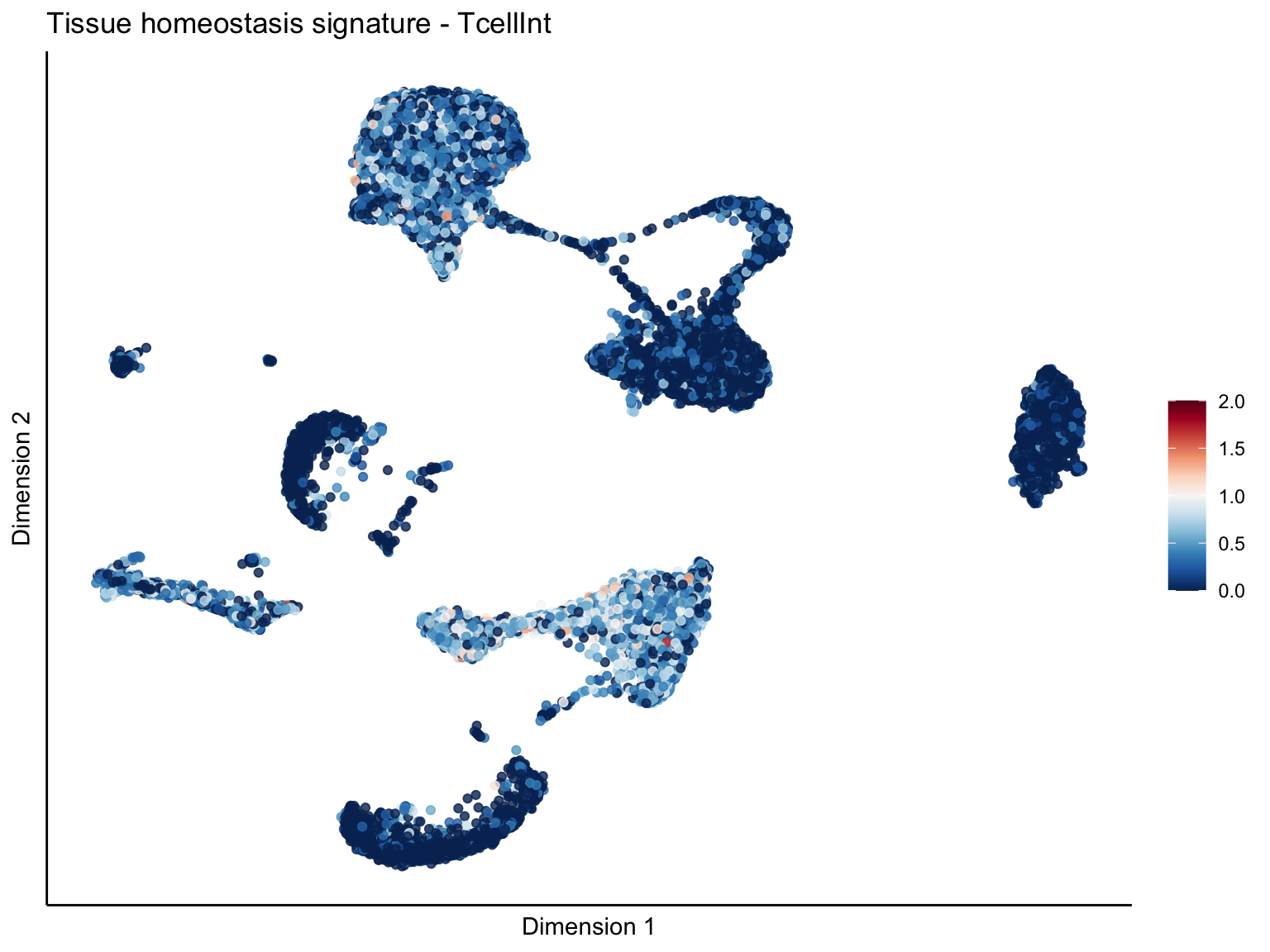
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]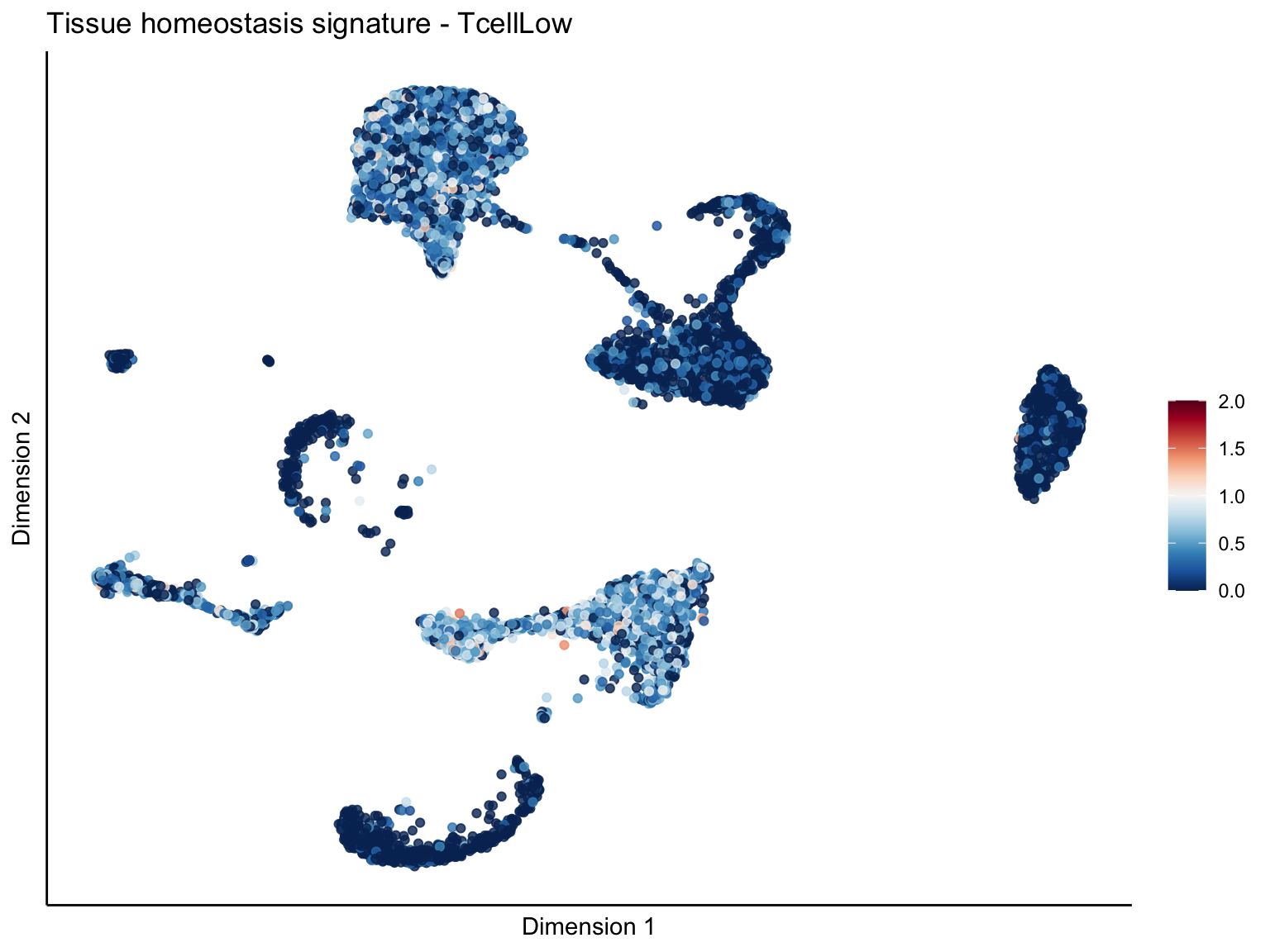
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
signature cut 1.5
cutOff <- 1.5
pal = colorRampPalette(rev(brewer.pal(11, 'RdBu')))
sc <- scale_colour_gradientn(colours = pal(100), limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]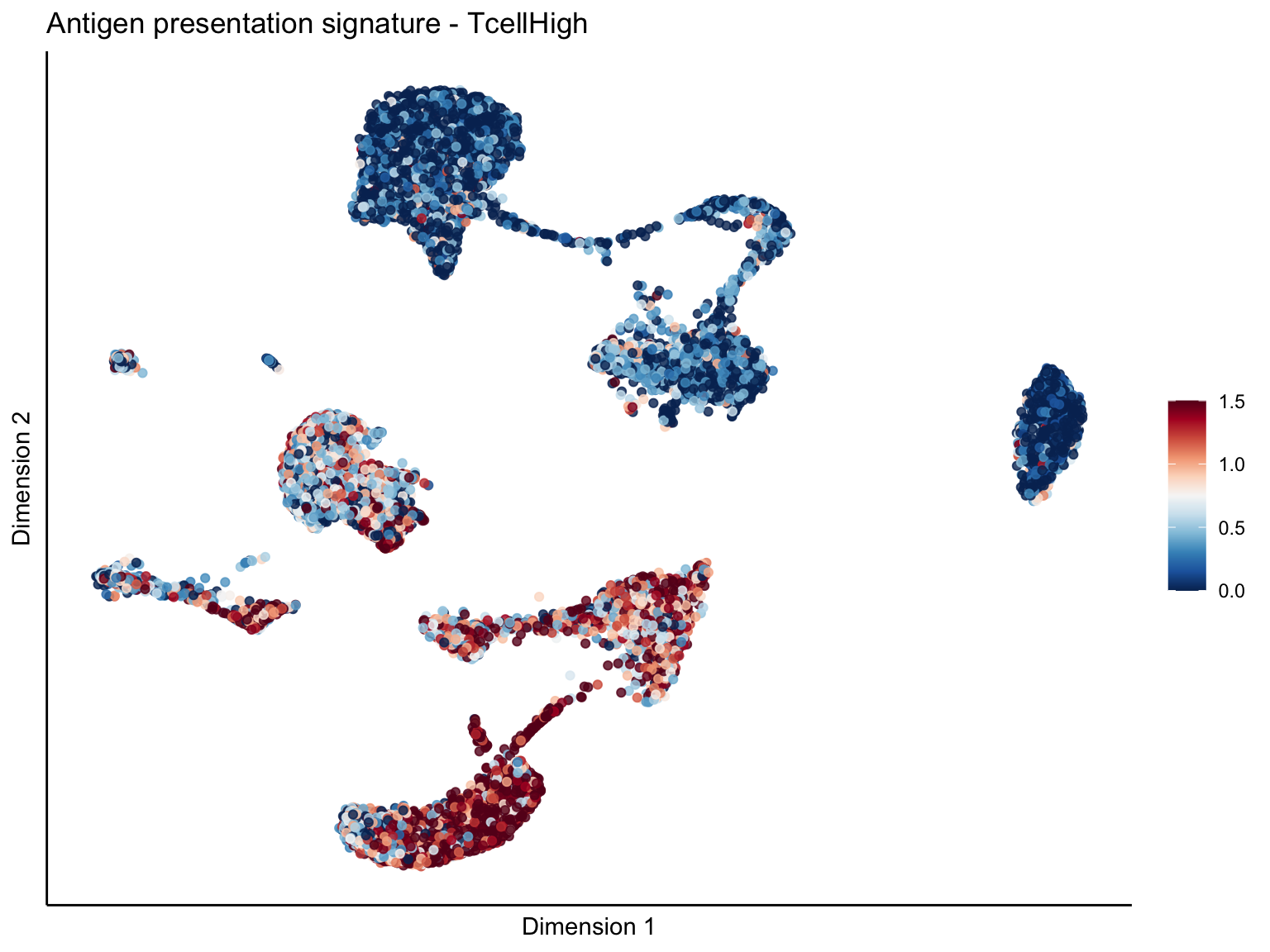
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]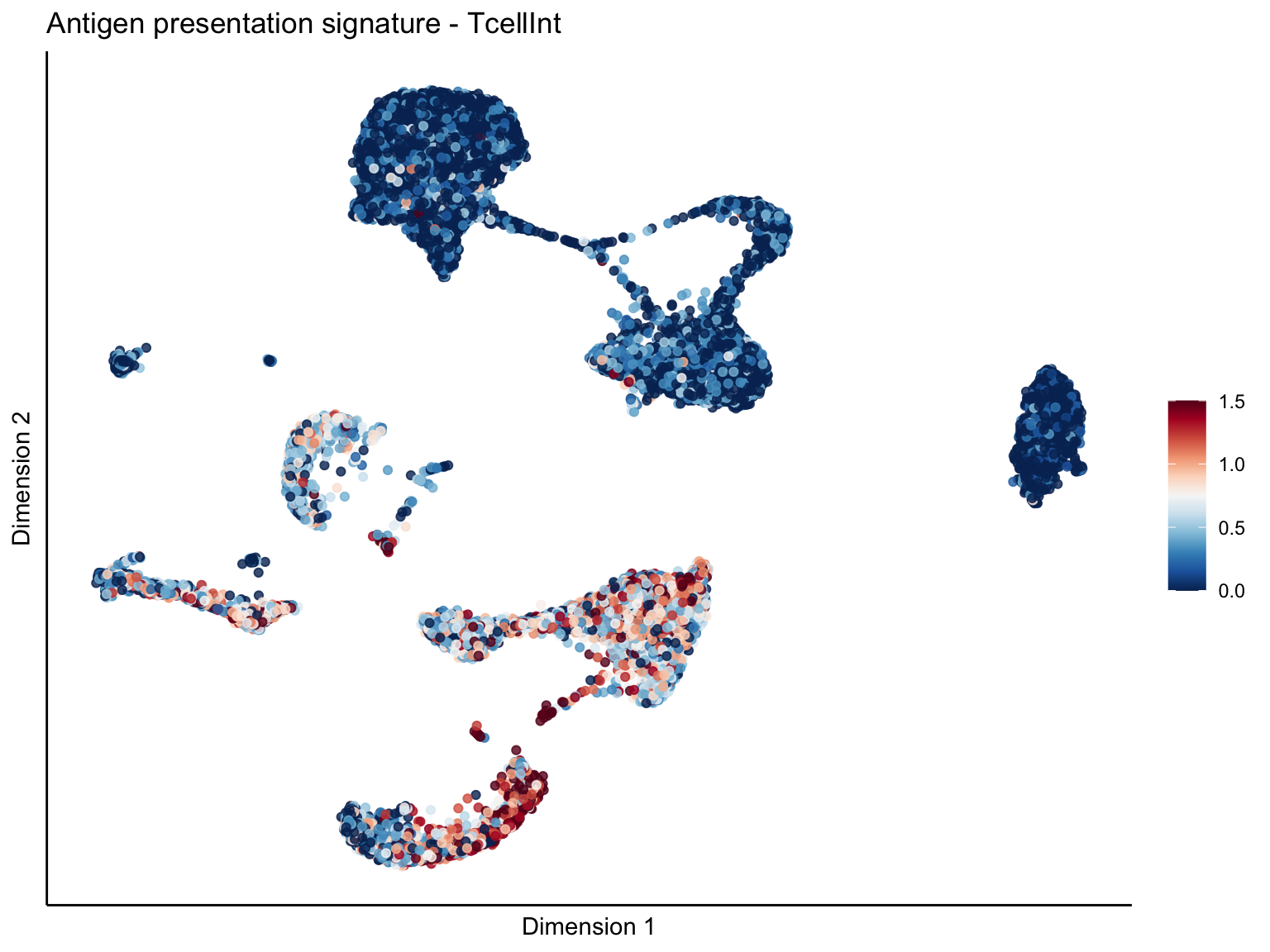
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]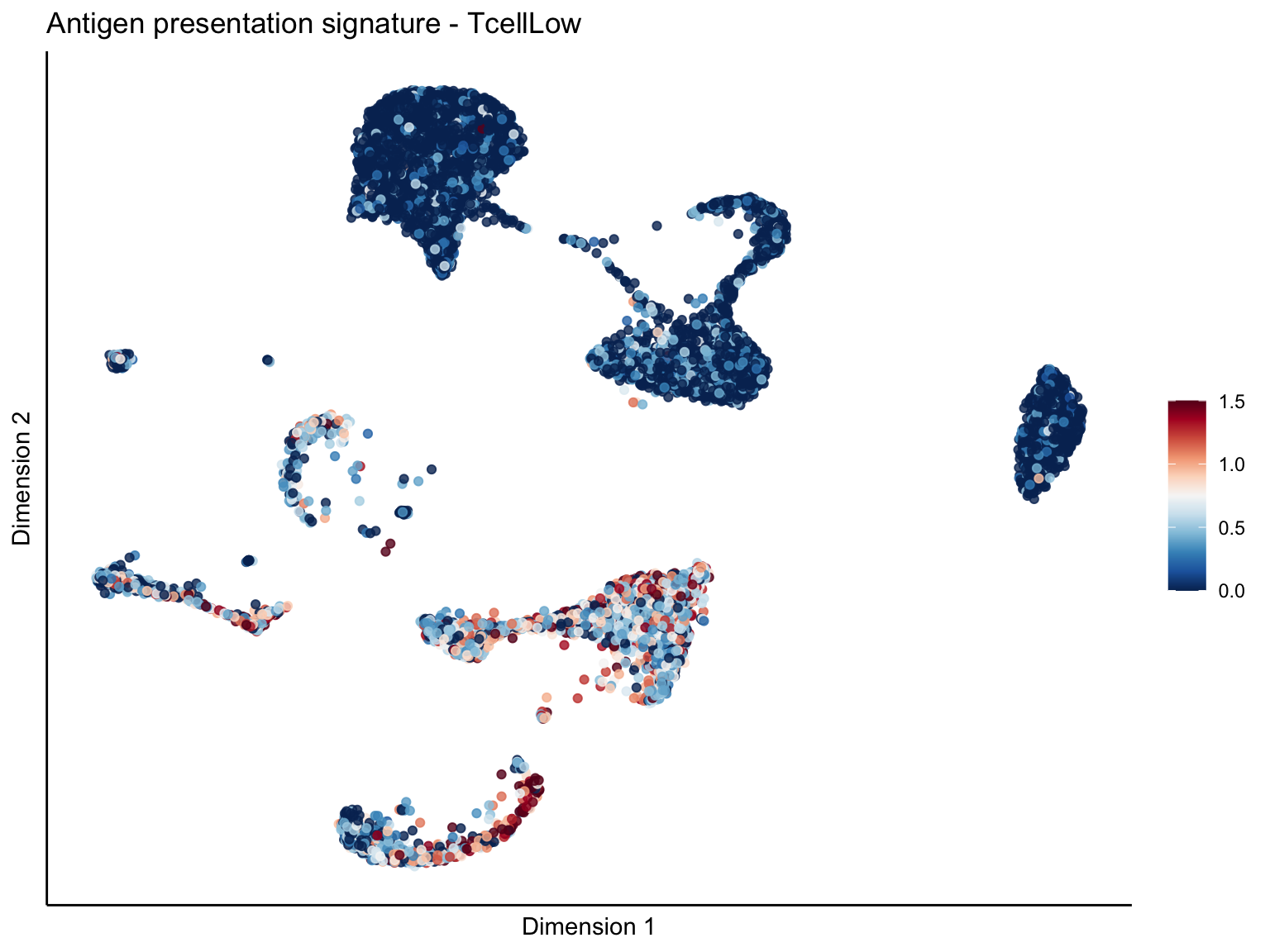
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]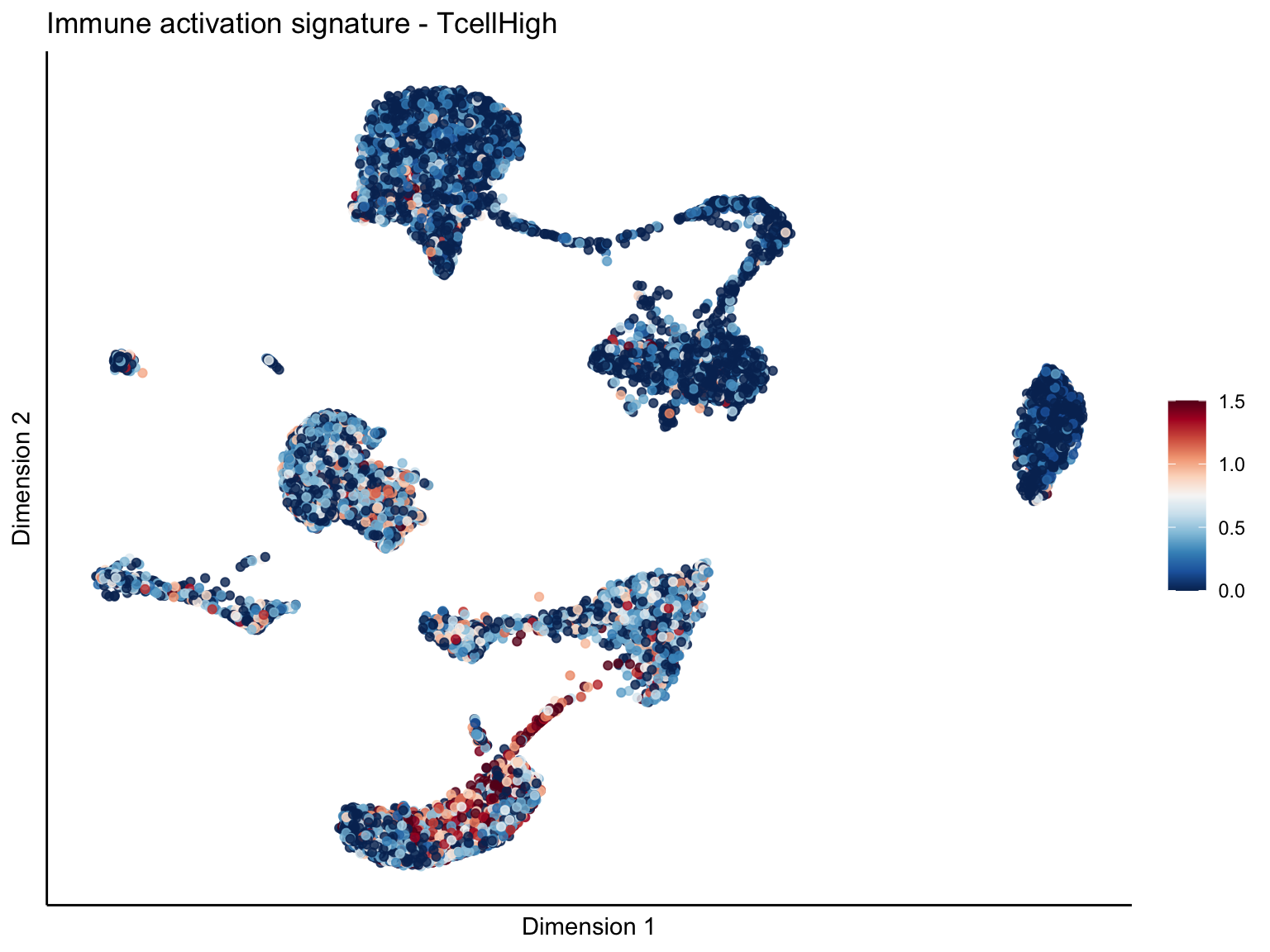
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]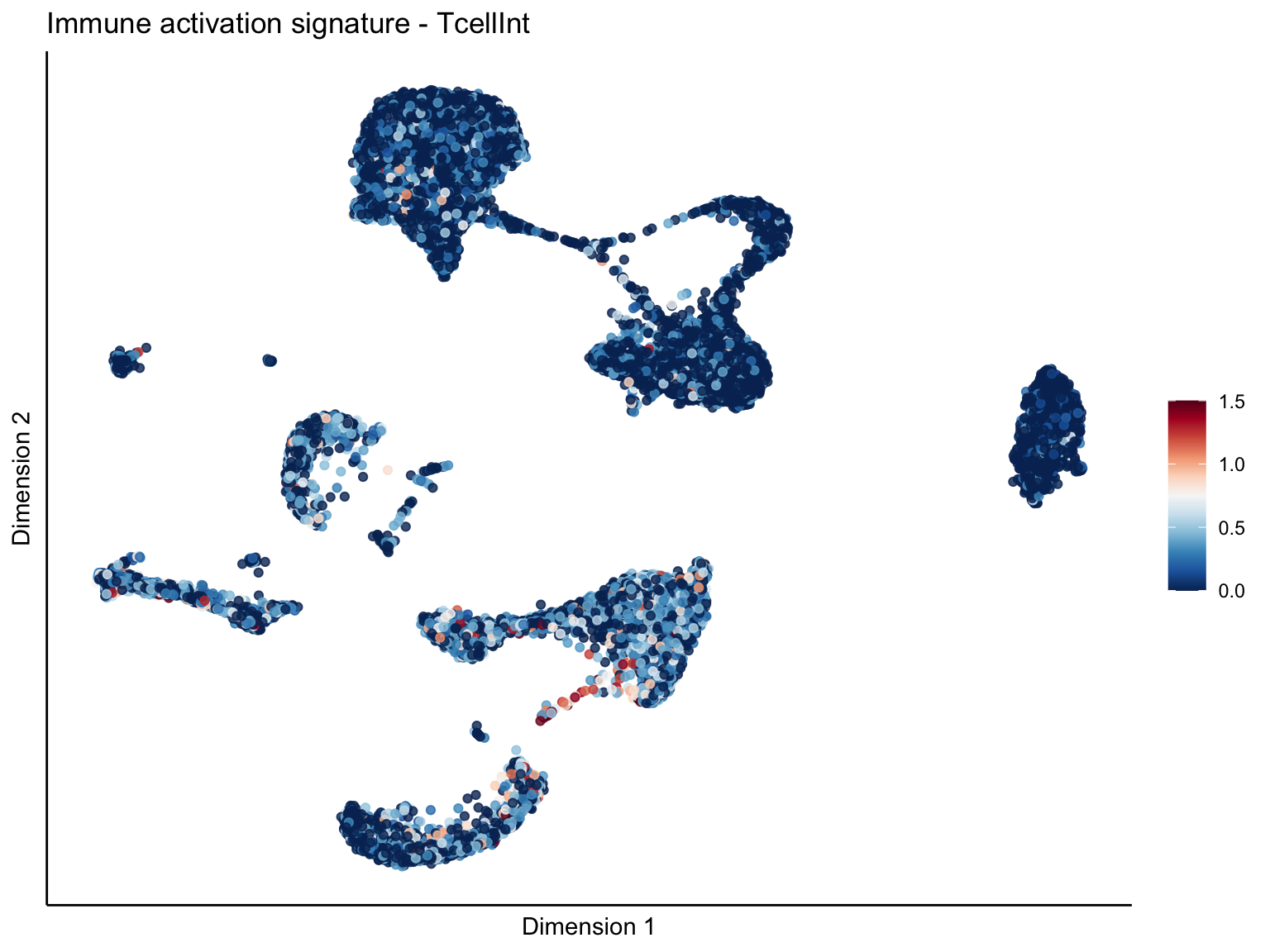
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]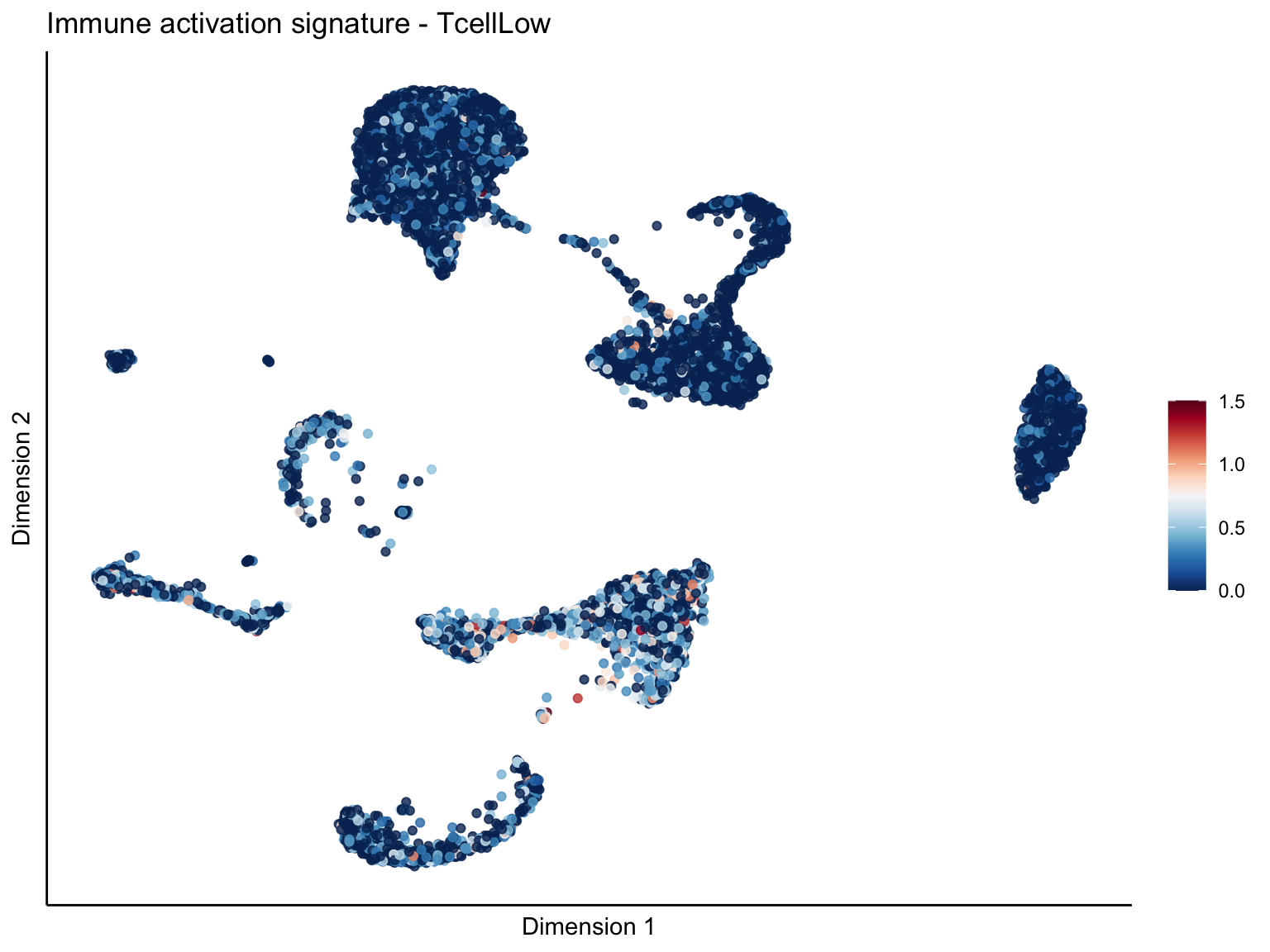
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]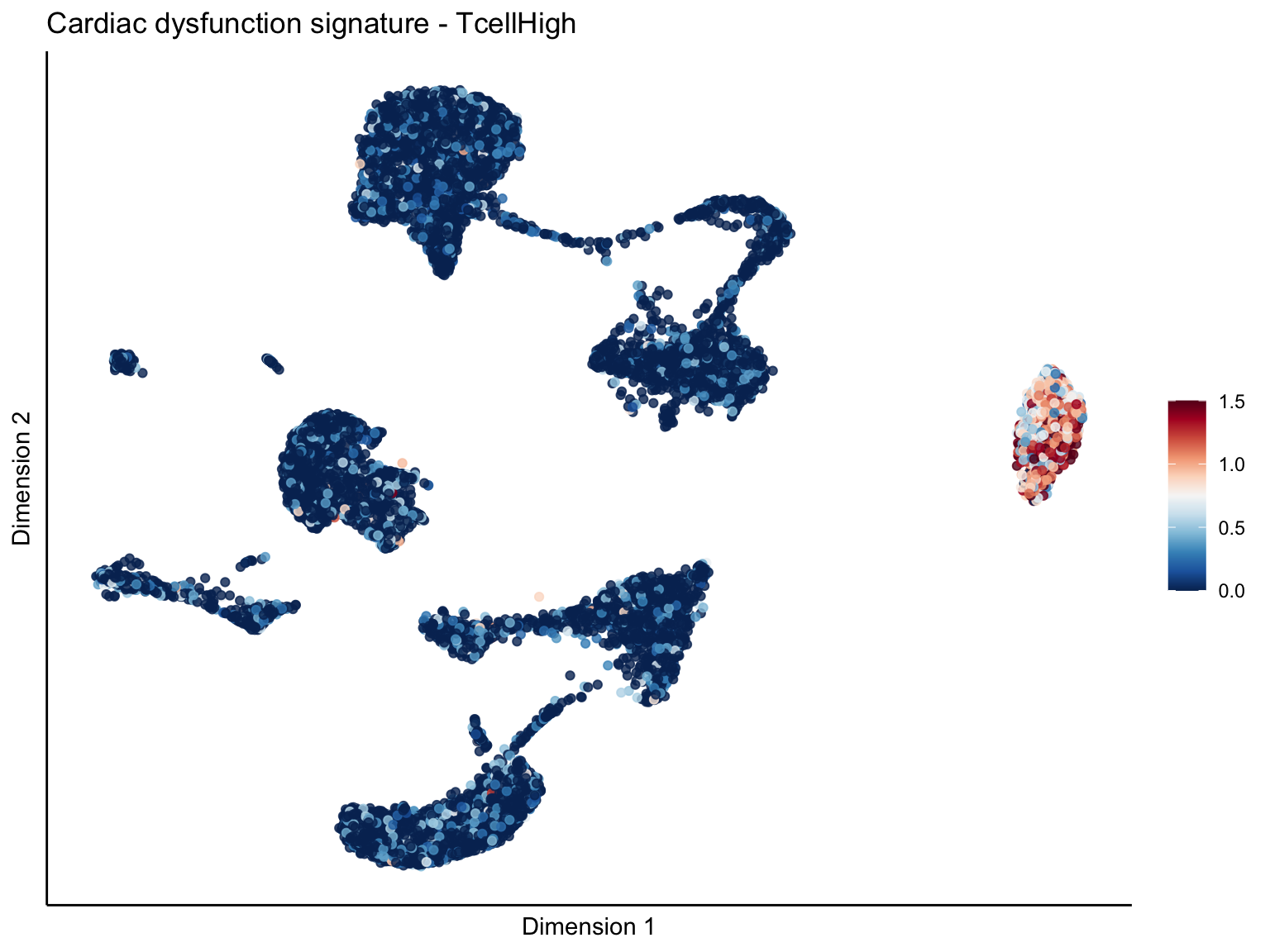
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]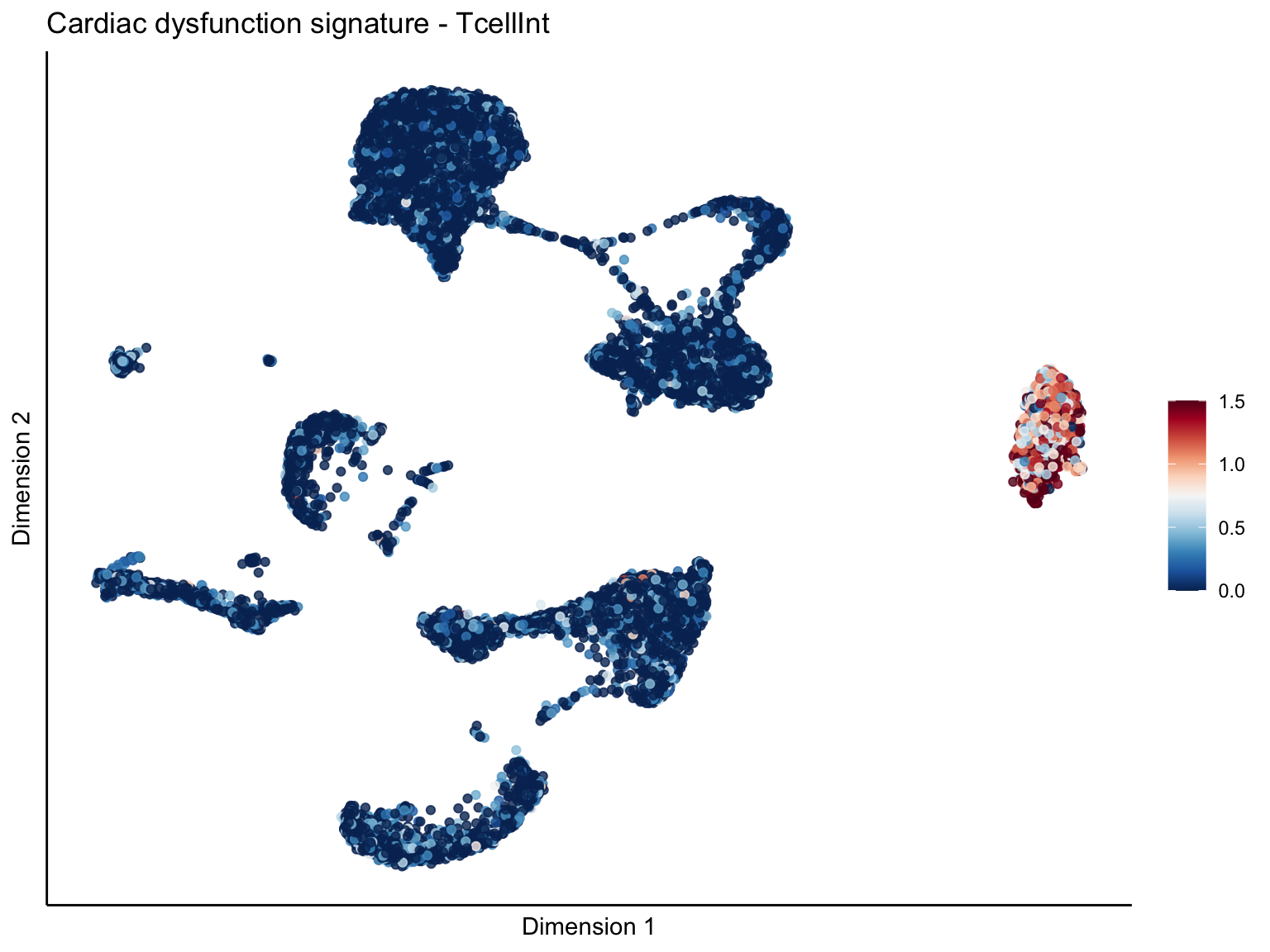
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]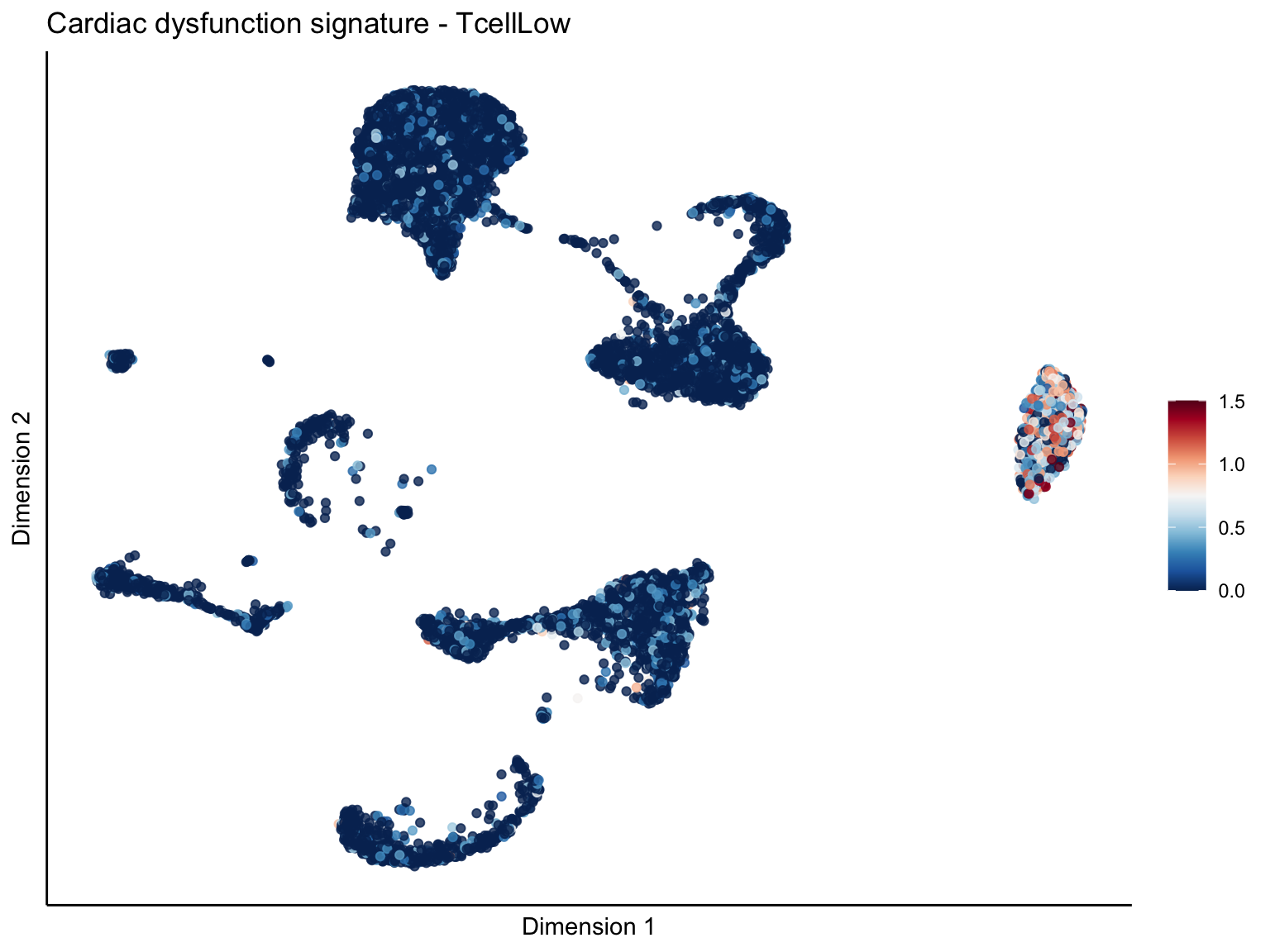
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]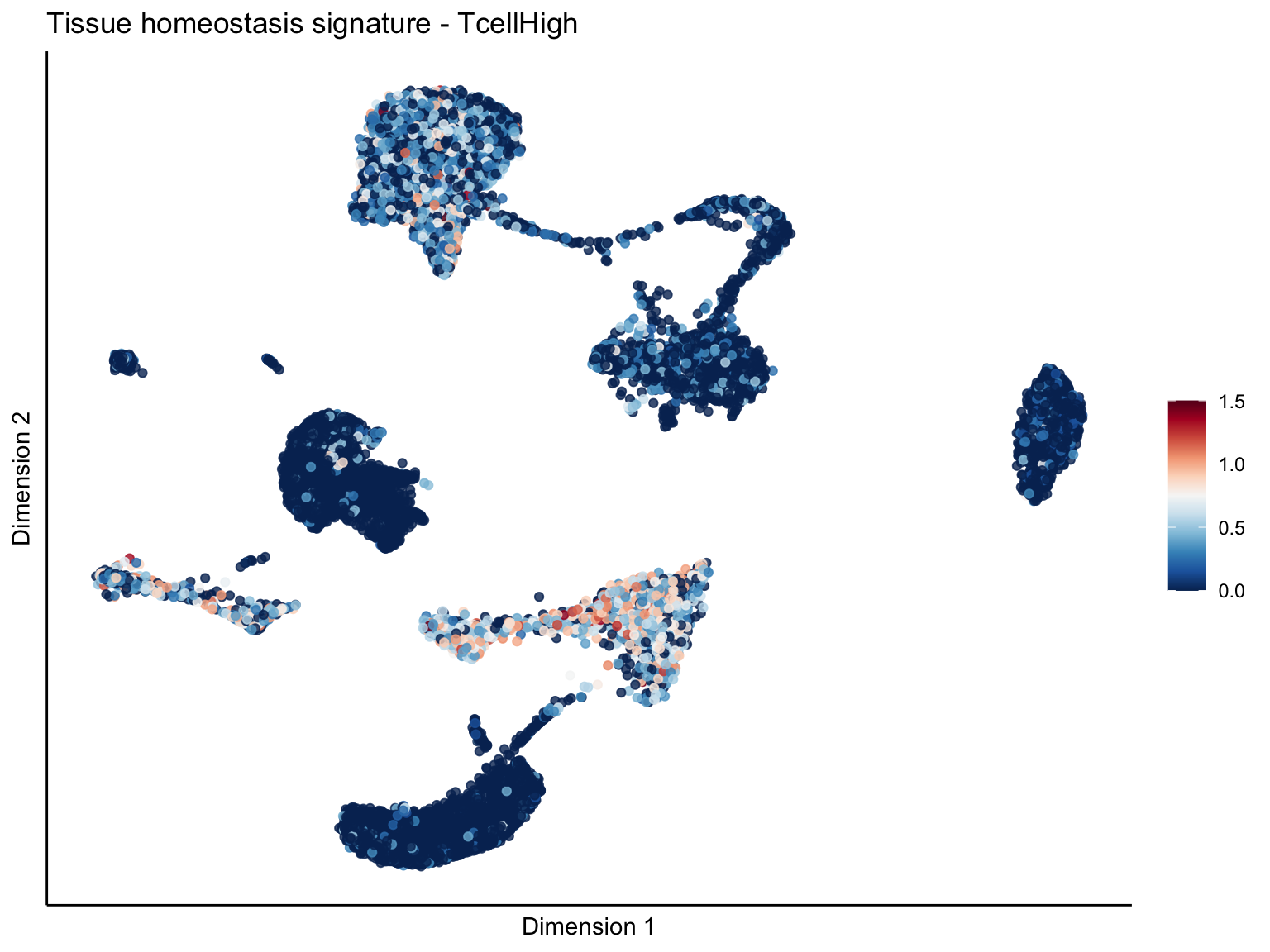
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]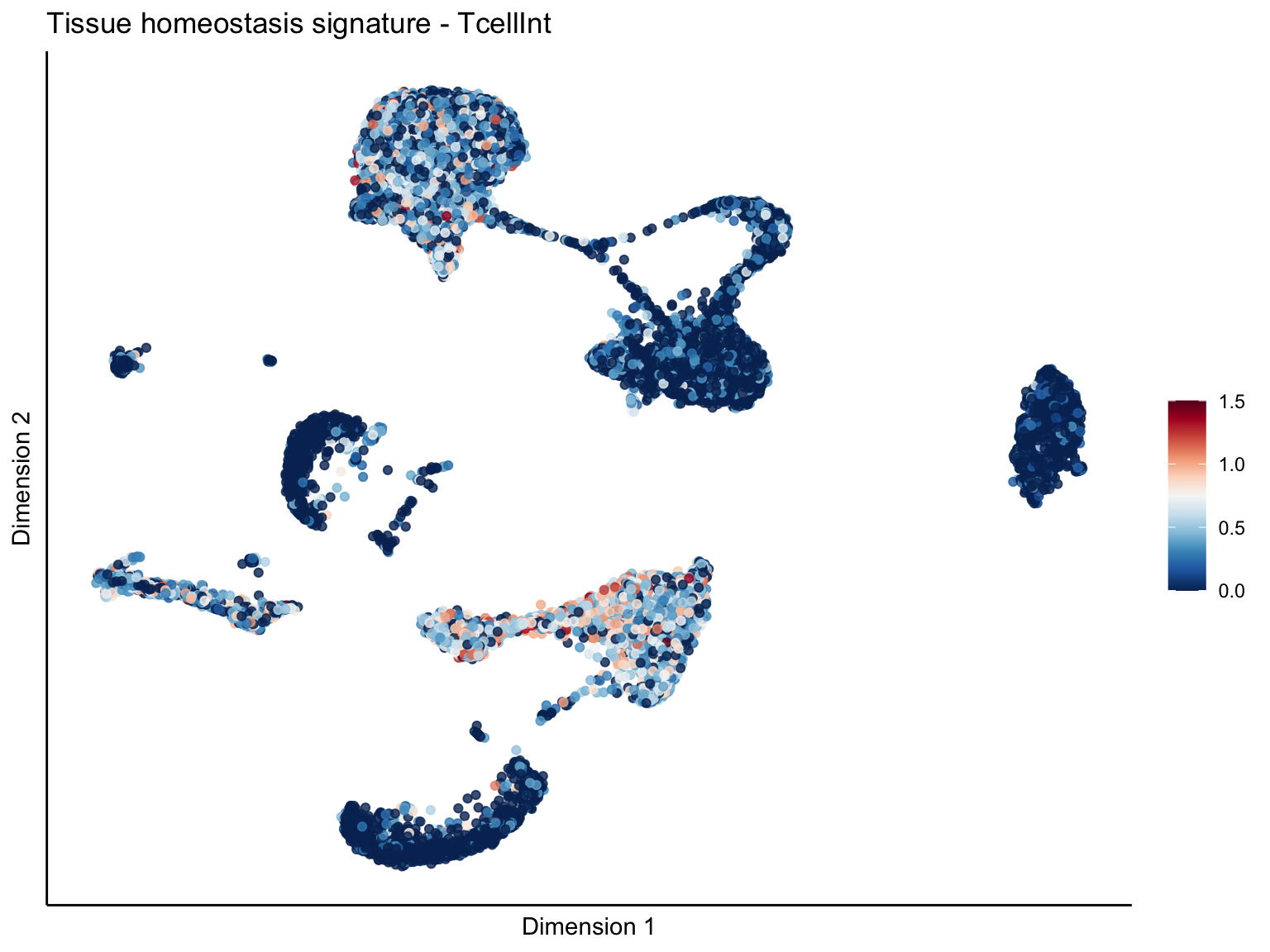
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]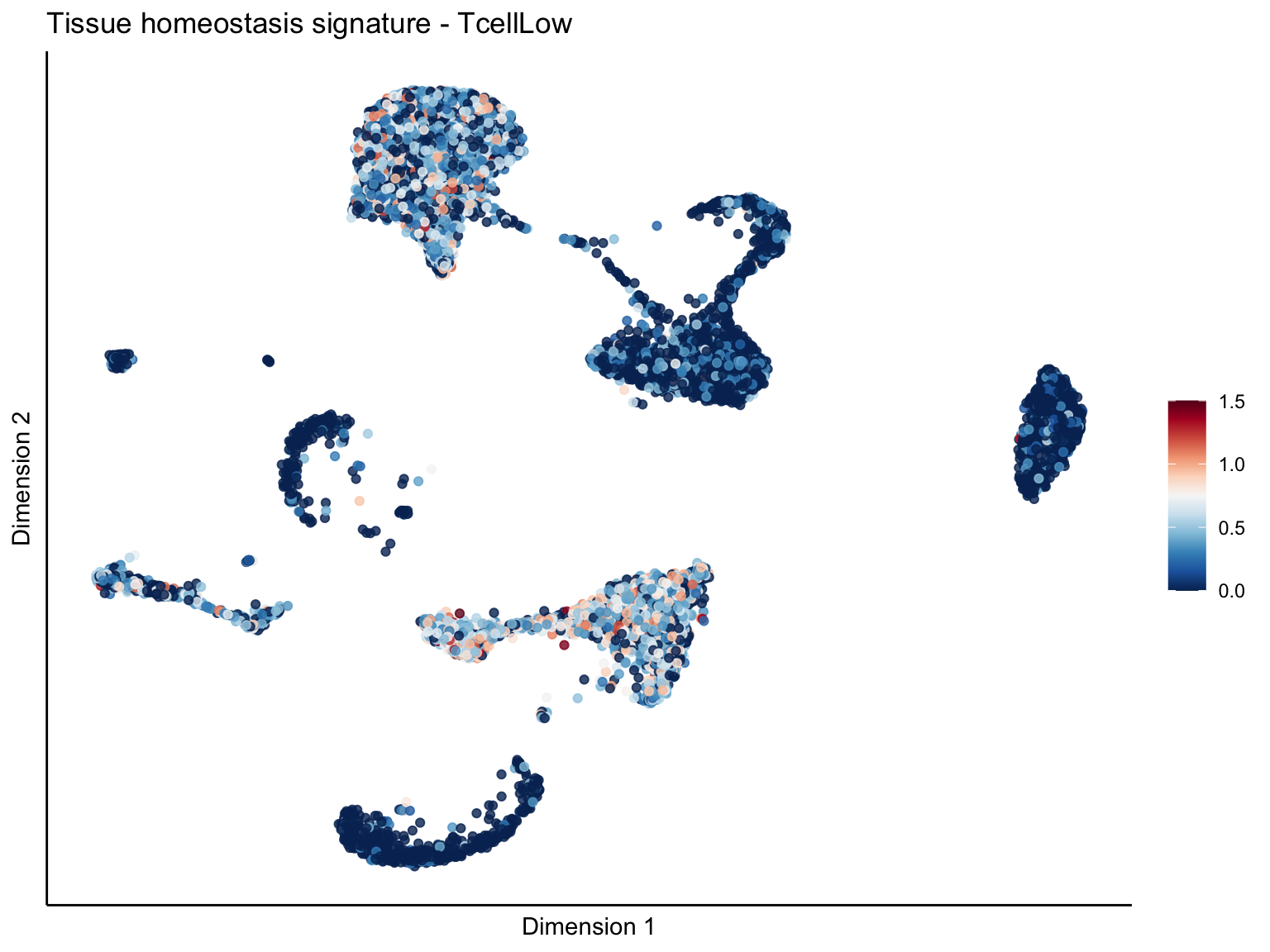
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
pal = viridis(100)
sc <- scale_colour_gradientn(colours = pal, limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]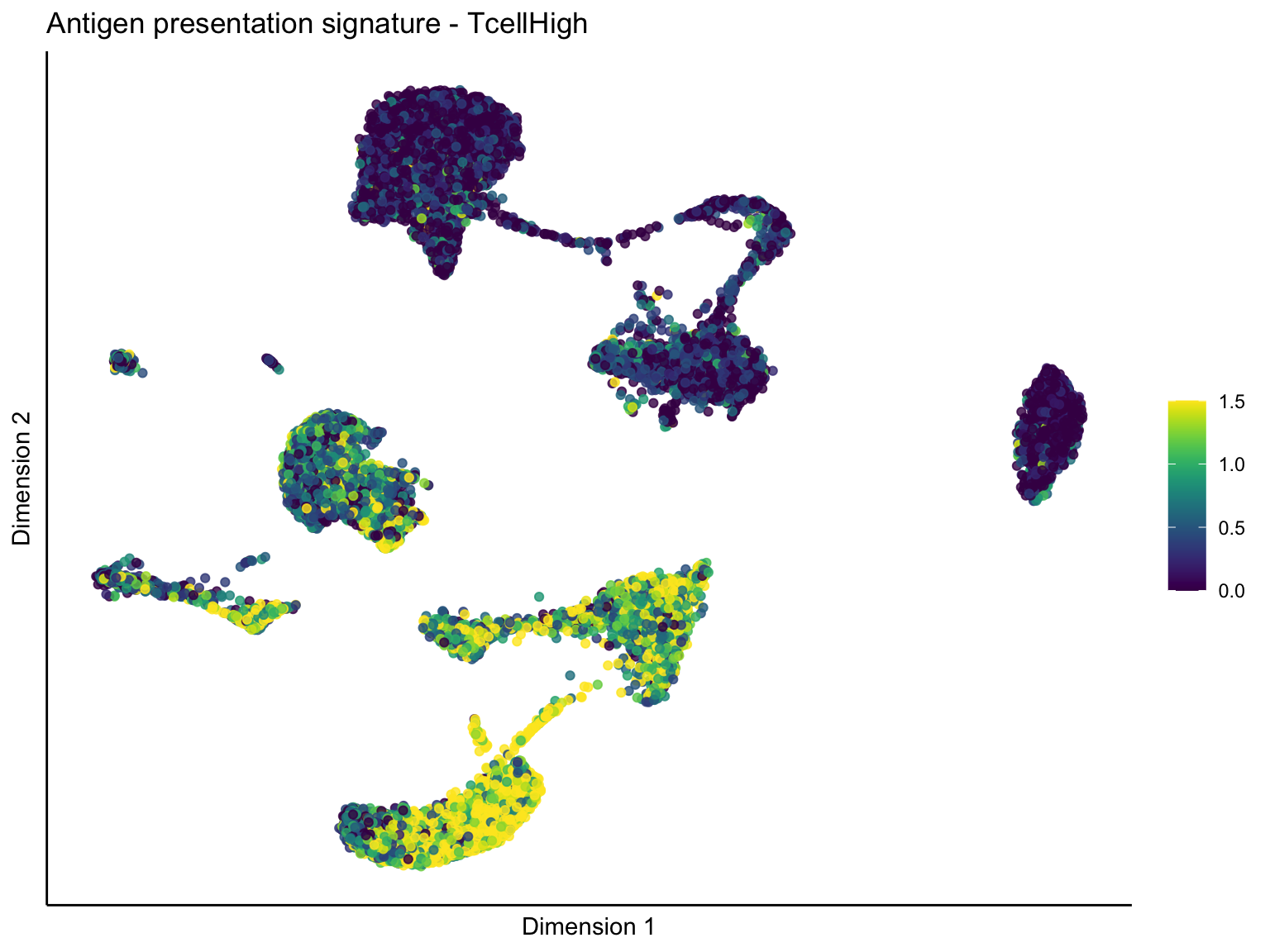
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]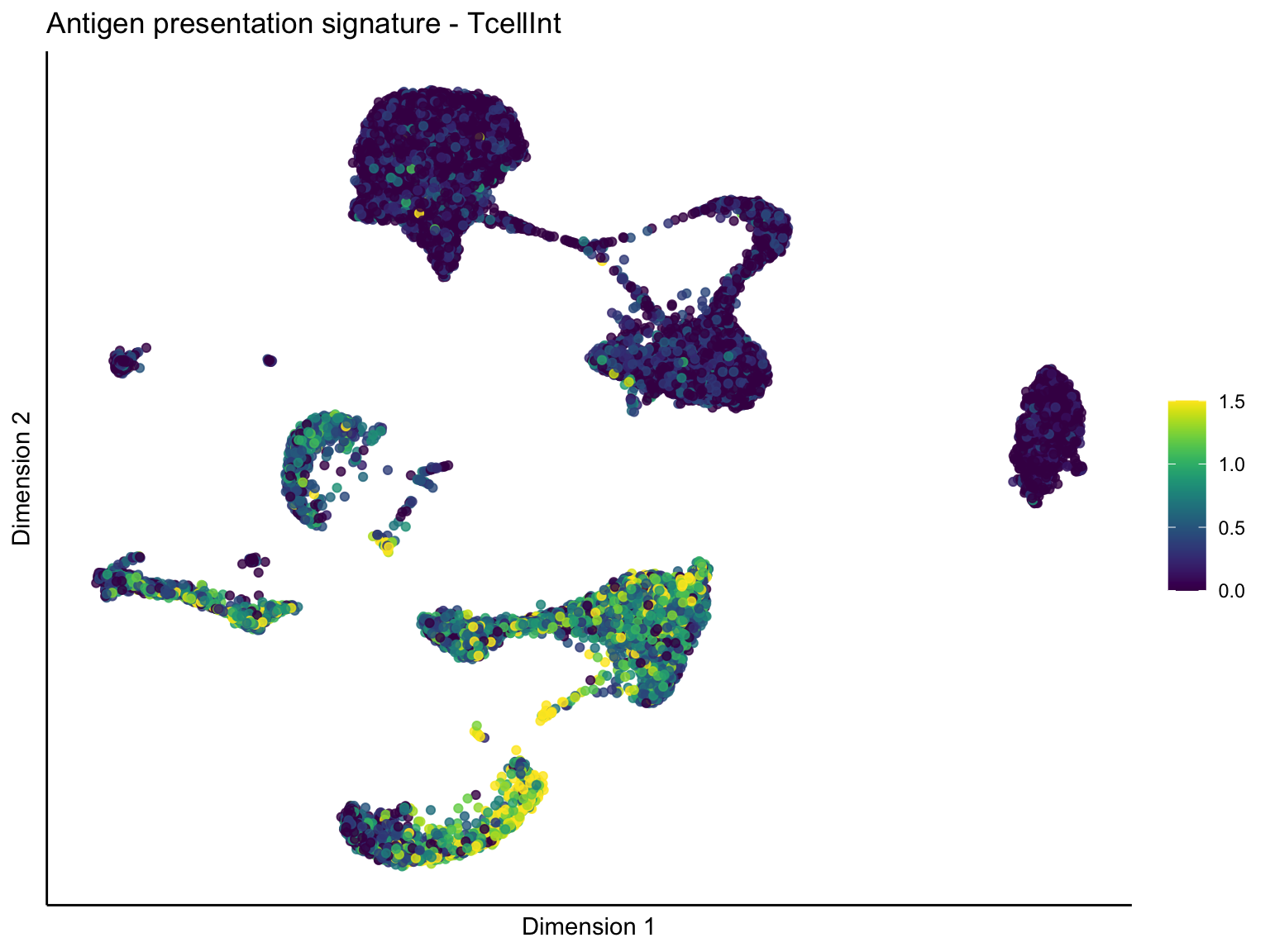
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]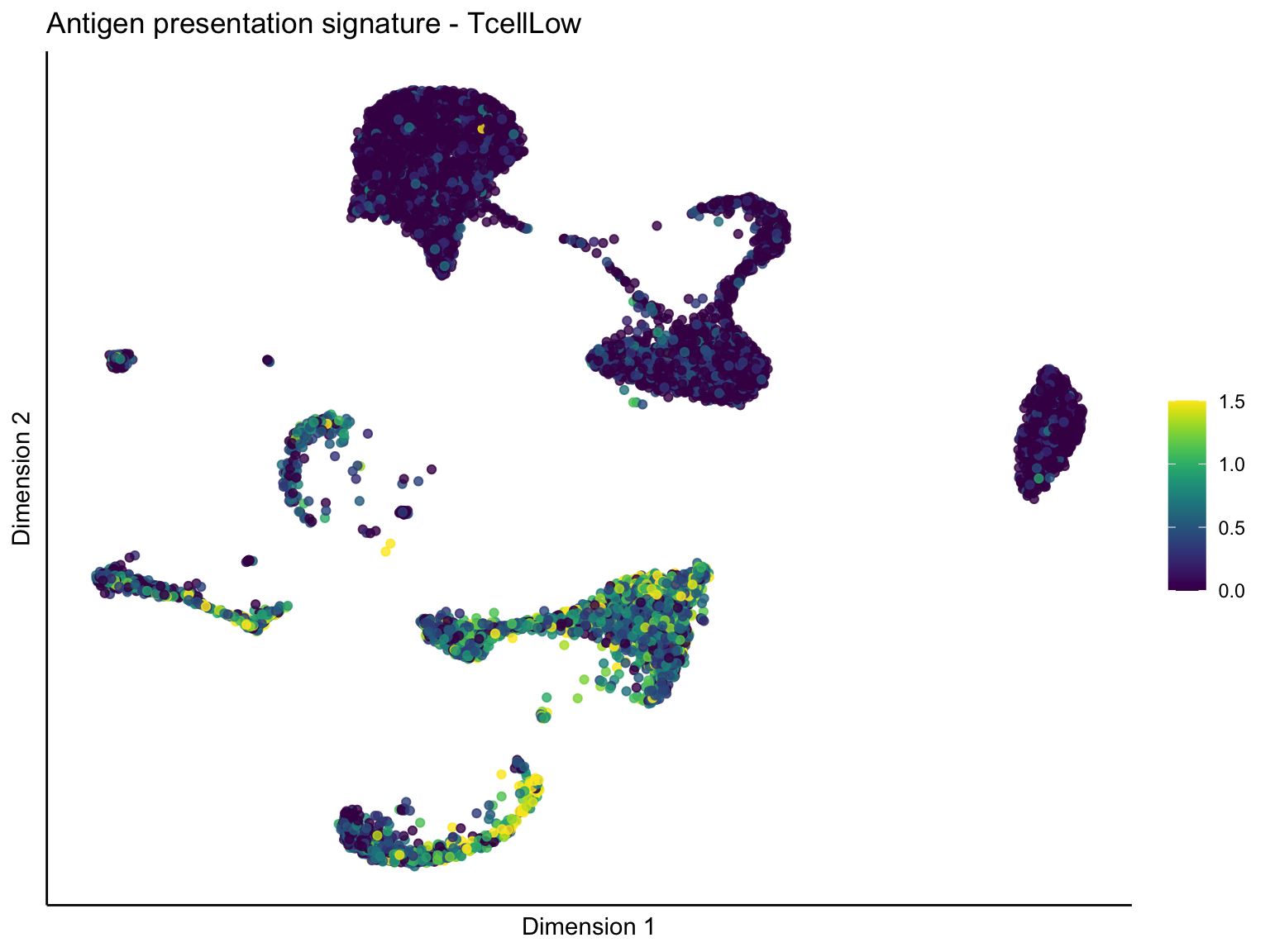
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]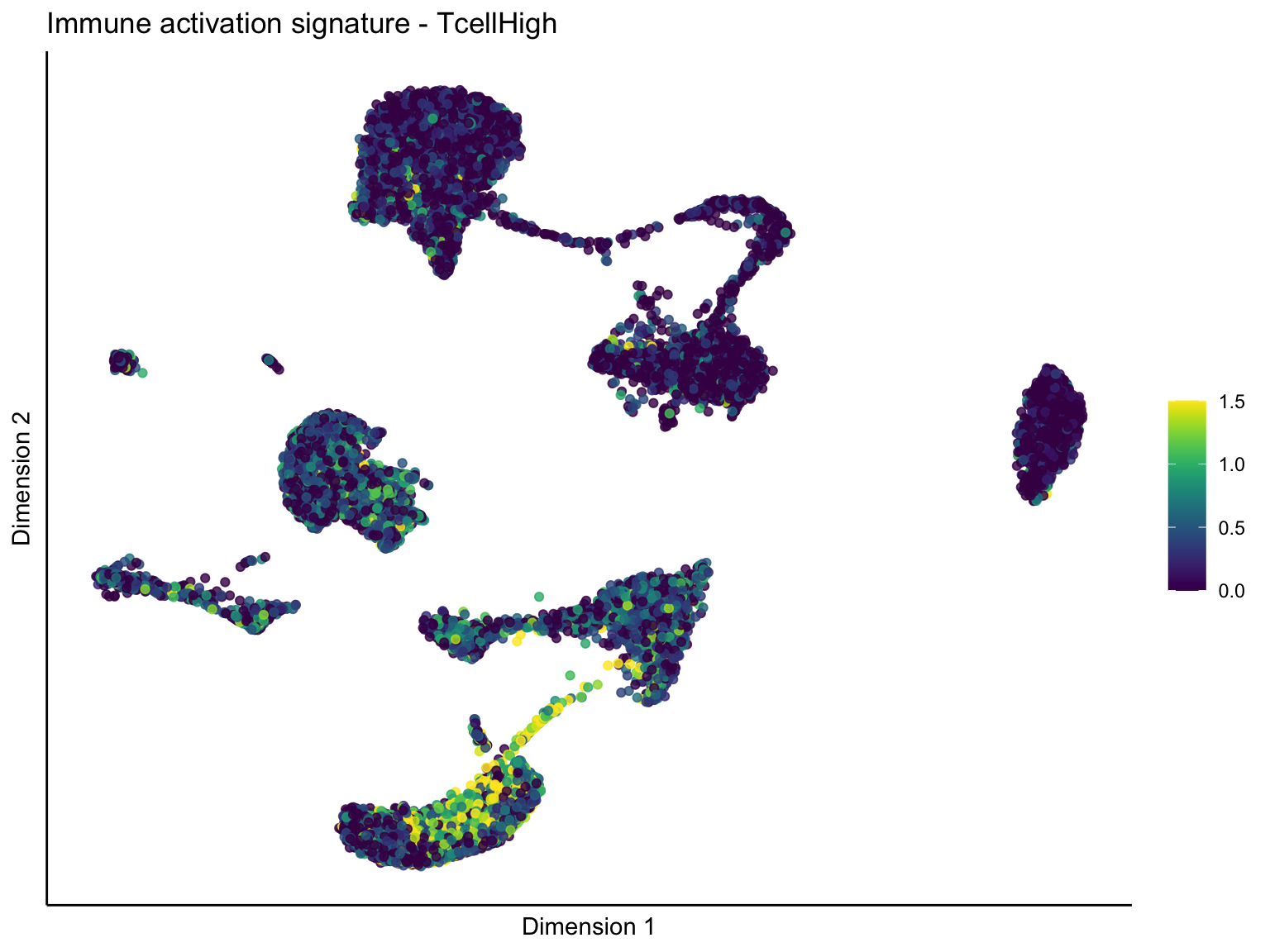
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]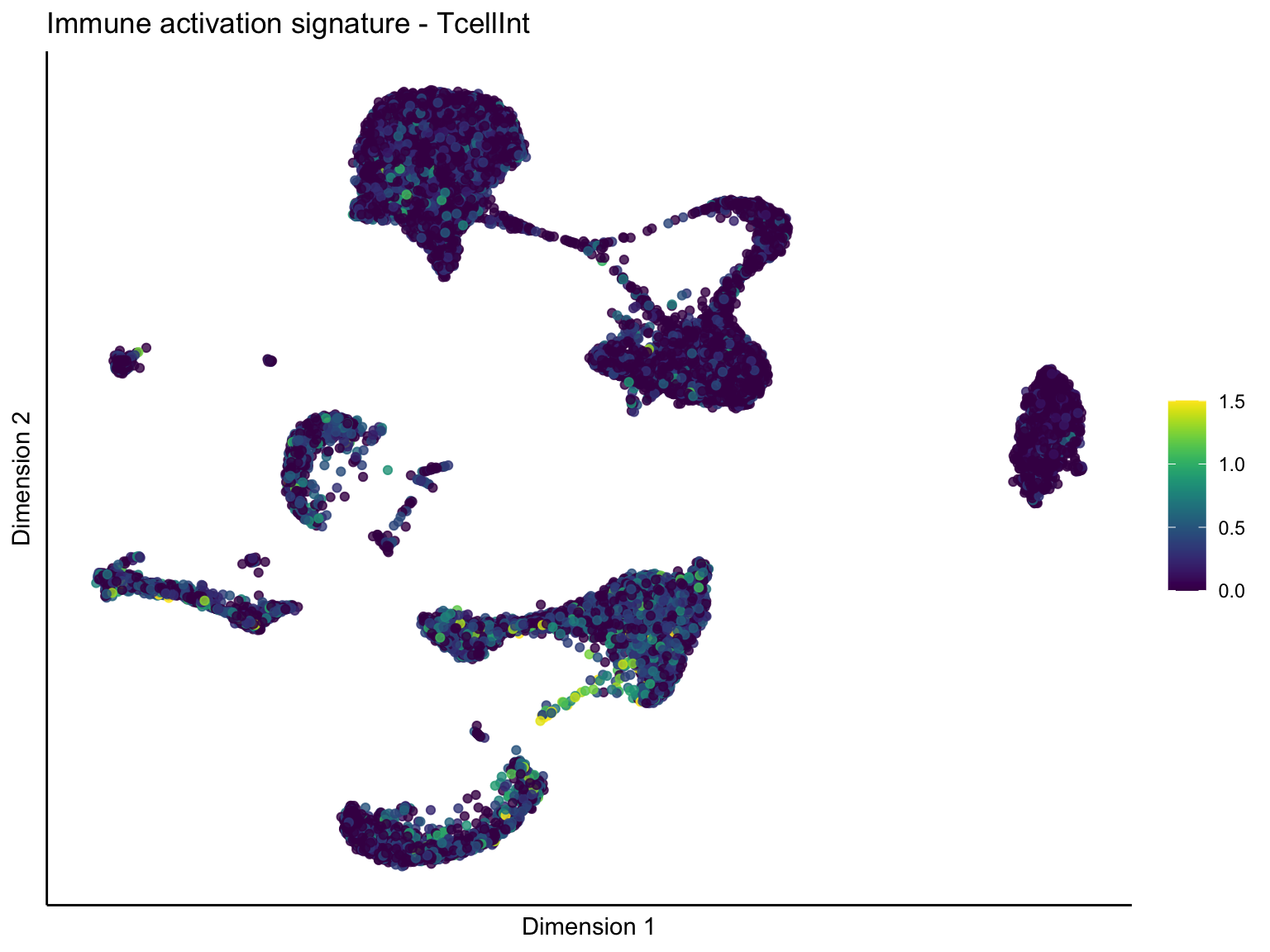
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]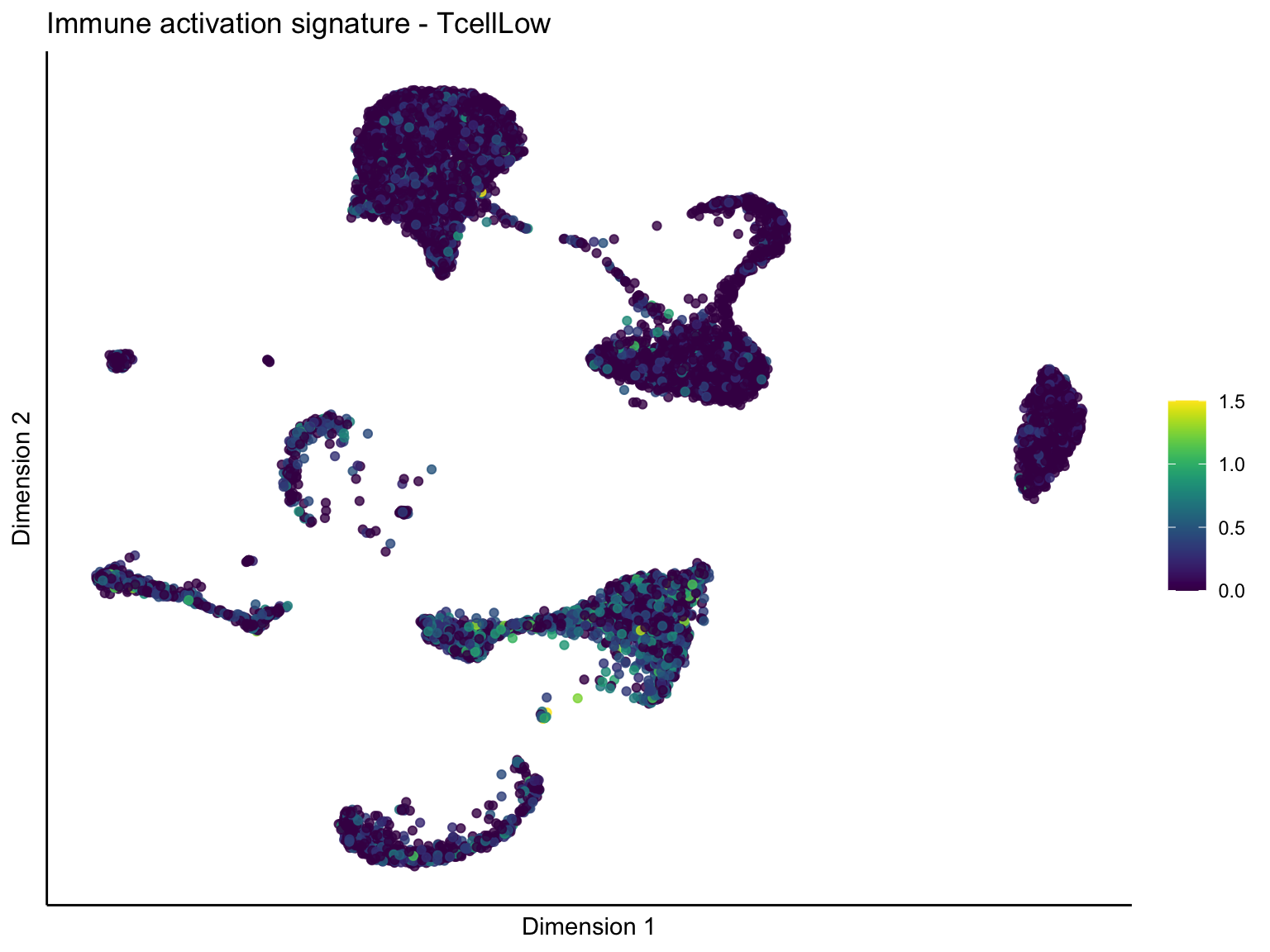
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]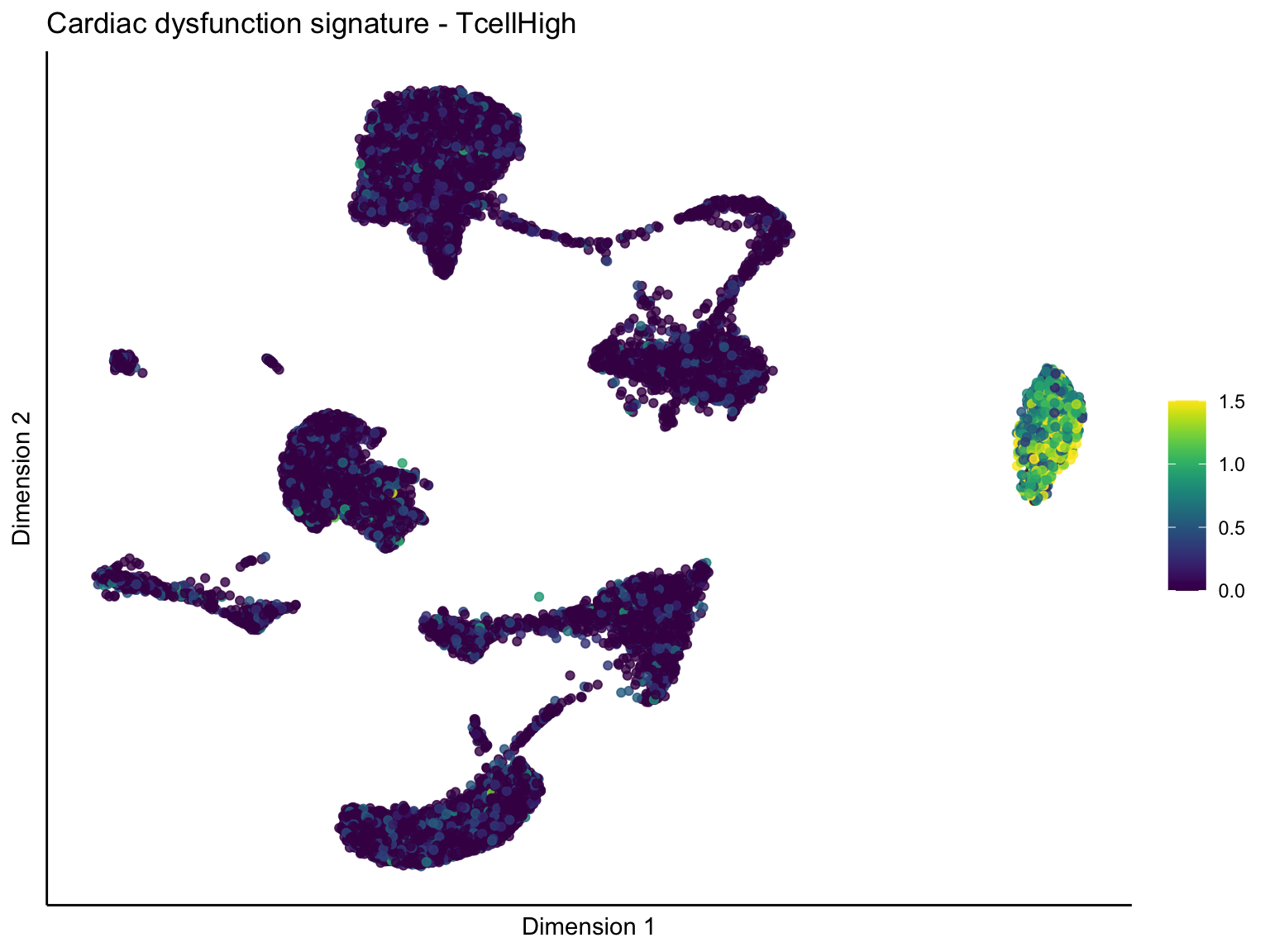
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]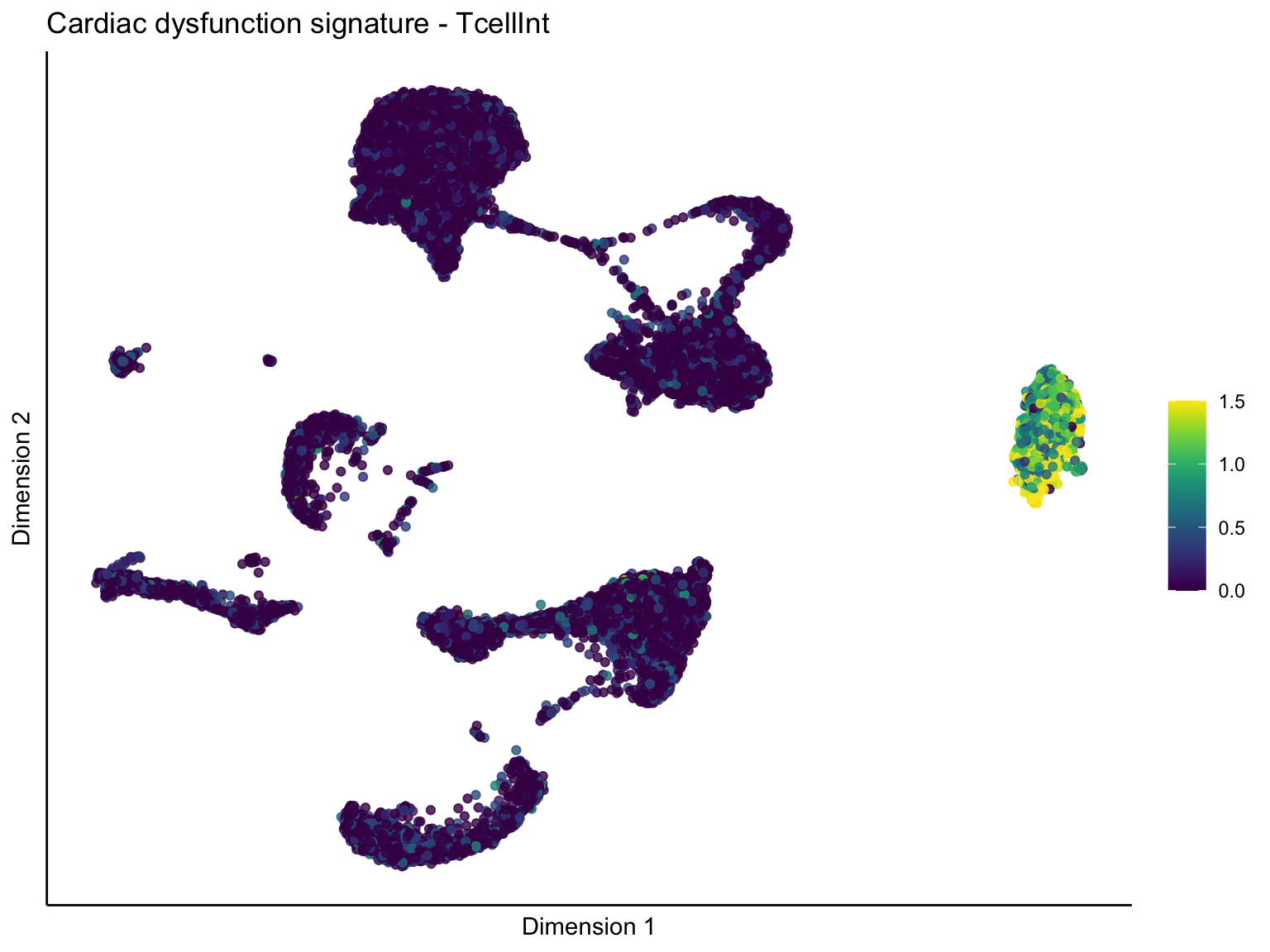
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]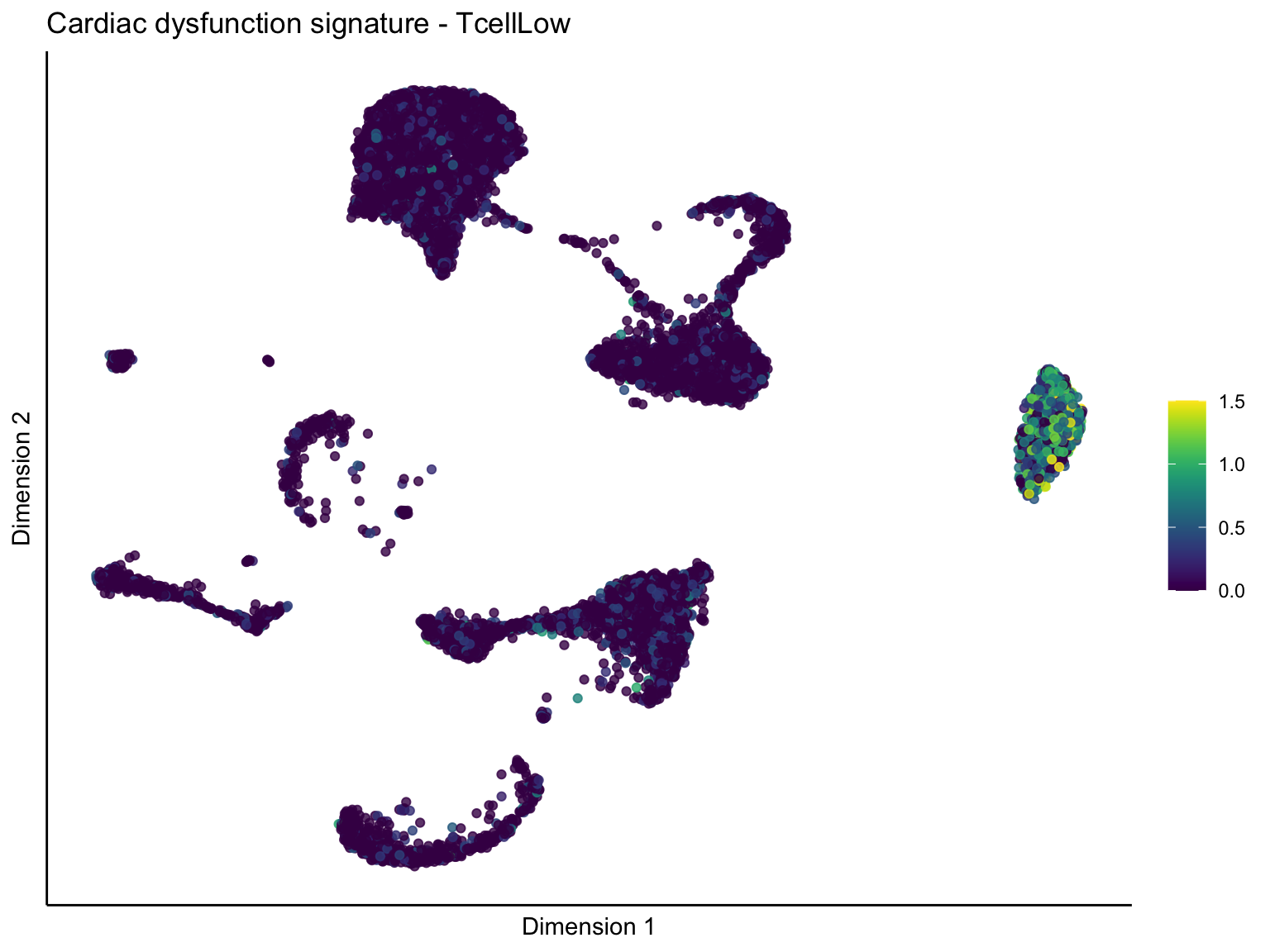
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]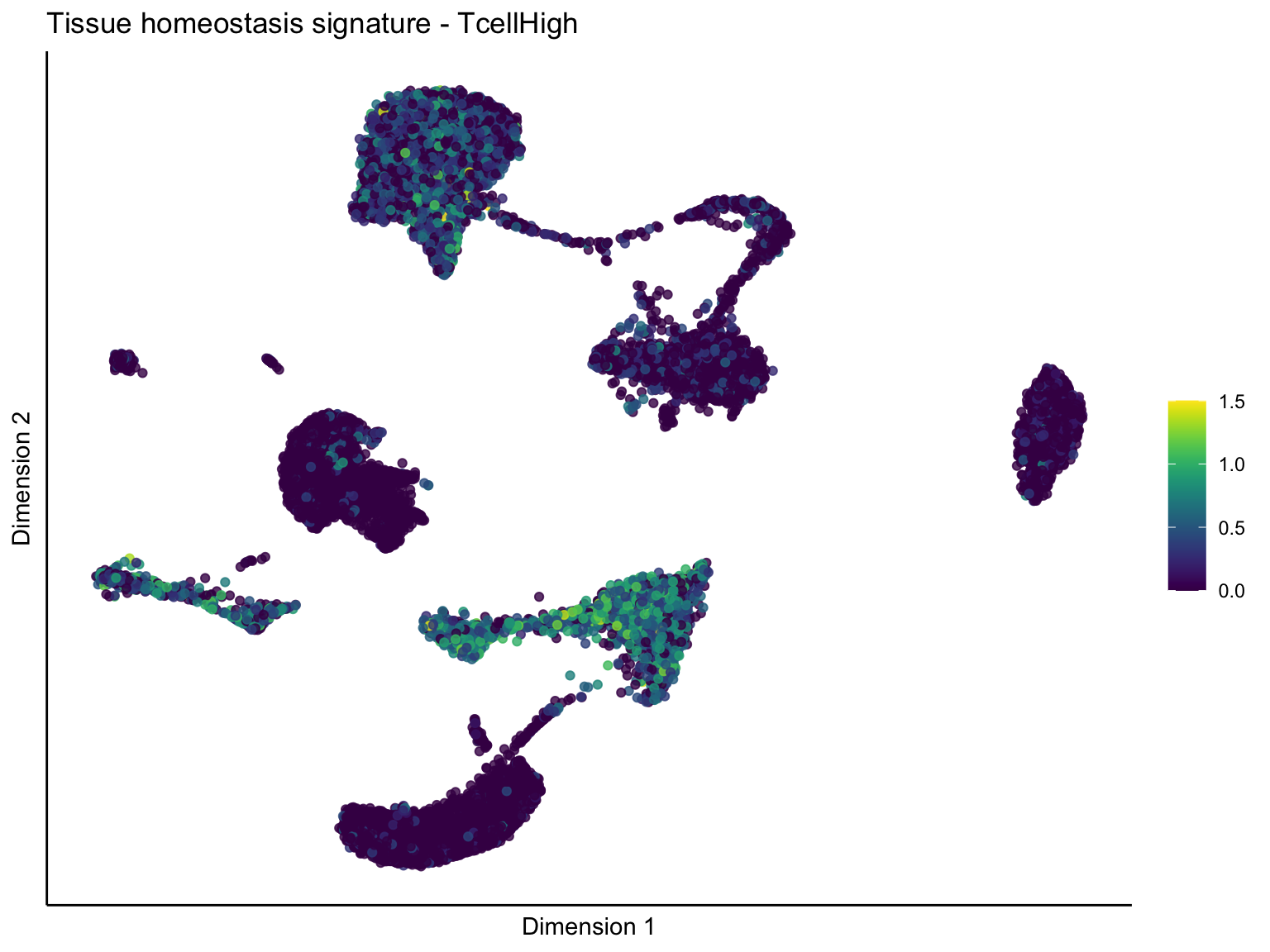
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]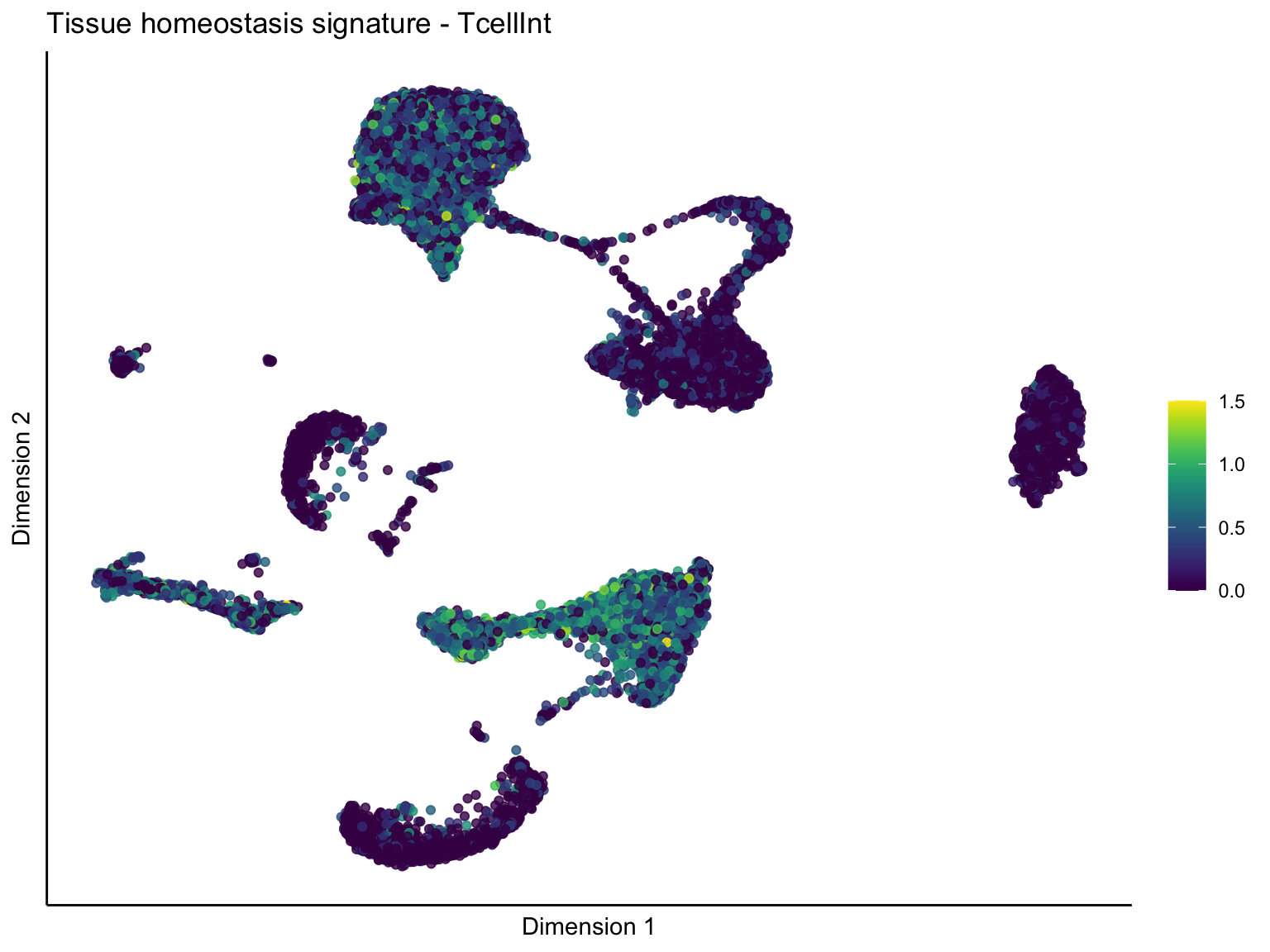
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]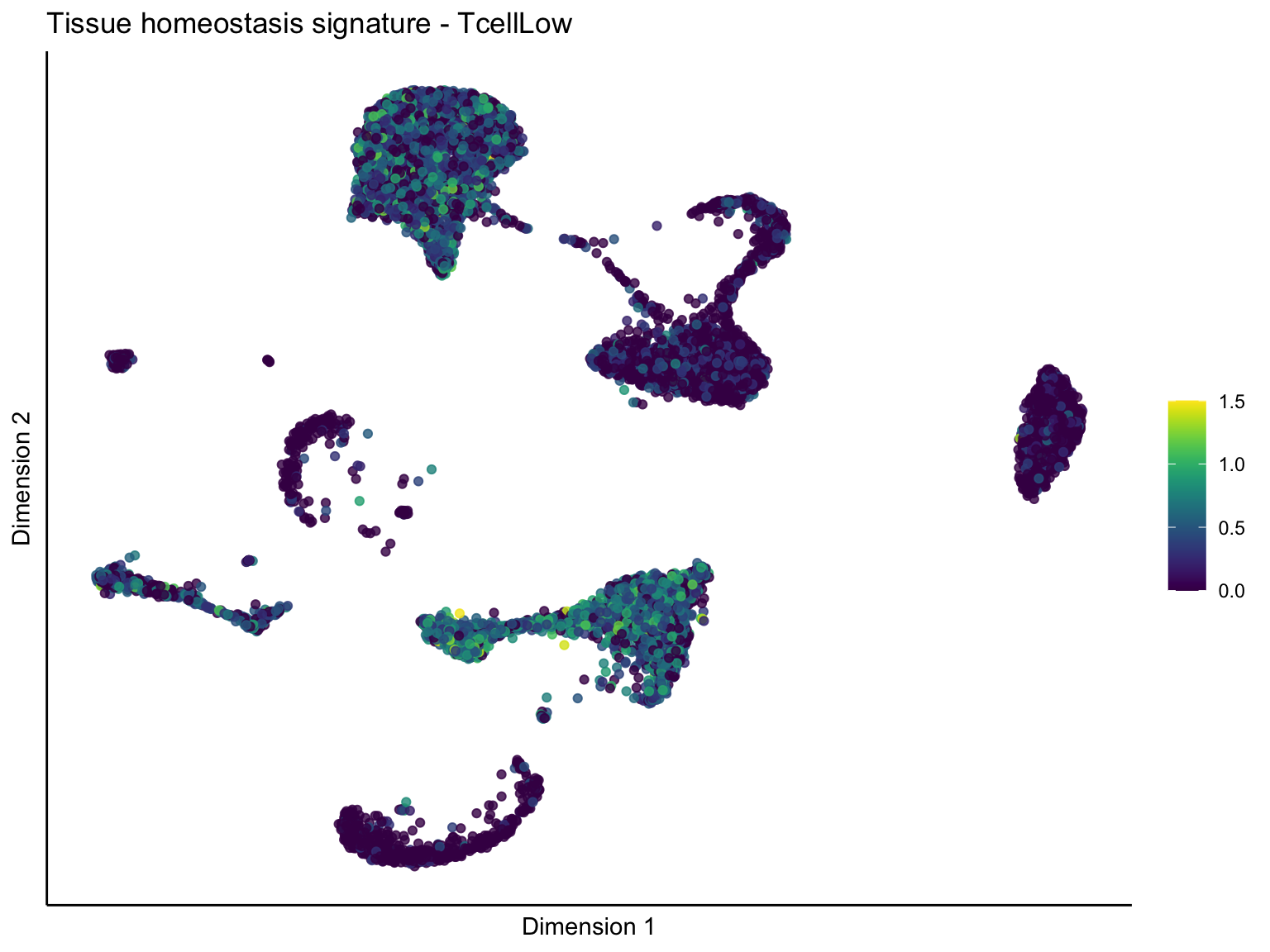
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
signature cut 1
cutOff <- 1
pal = colorRampPalette(rev(brewer.pal(11, 'RdBu')))
sc <- scale_colour_gradientn(colours = pal(100), limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]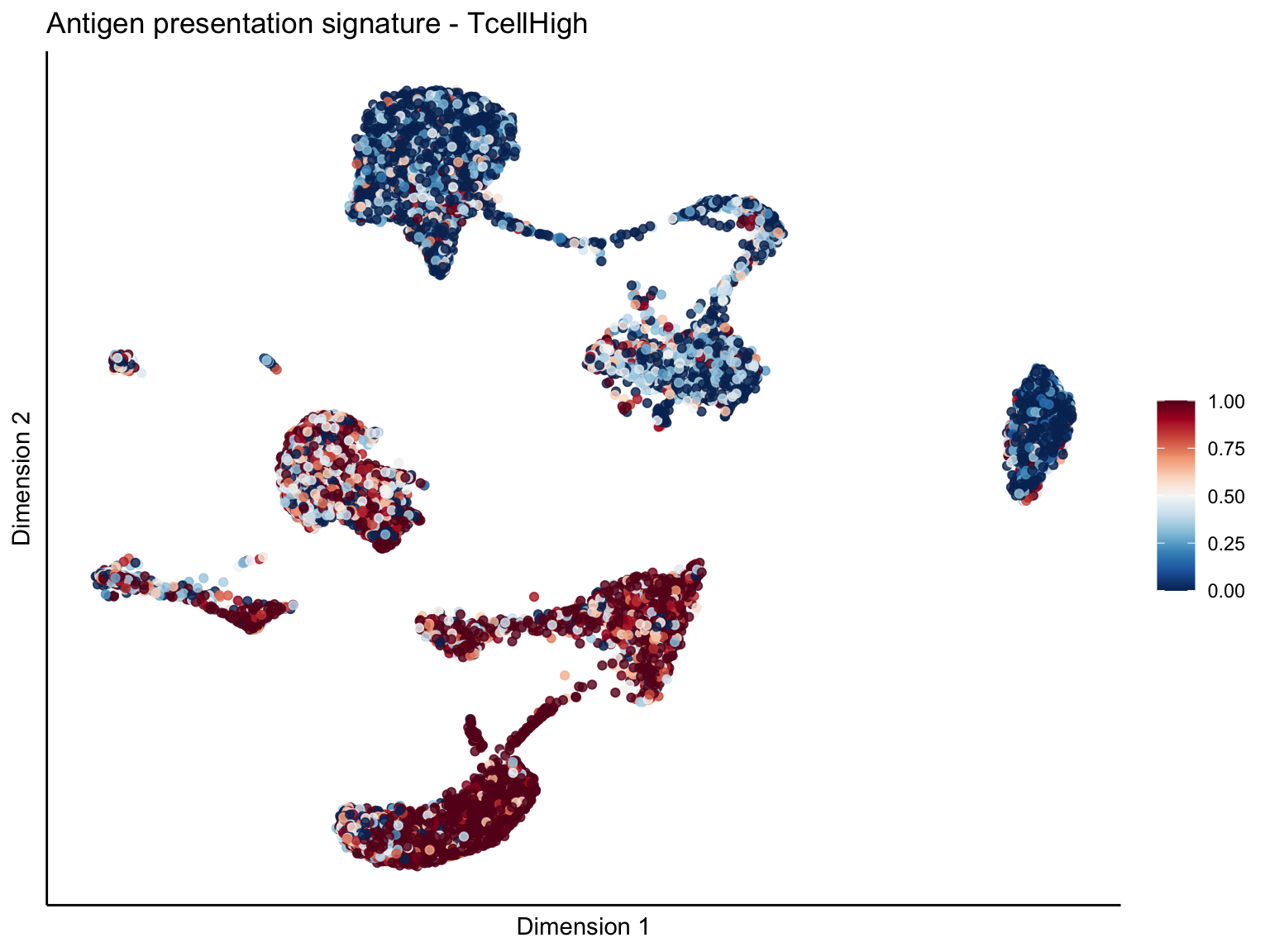
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]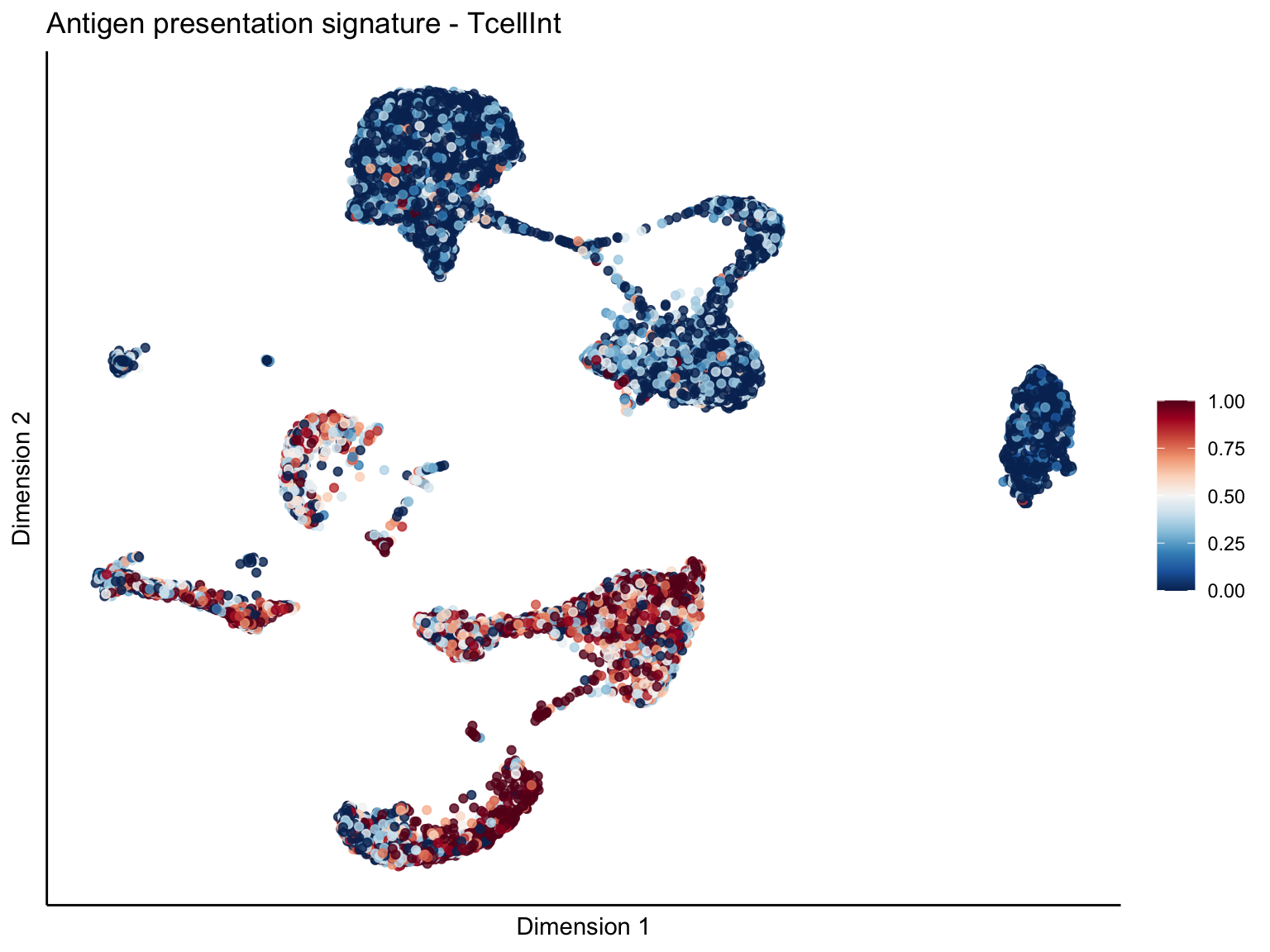
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]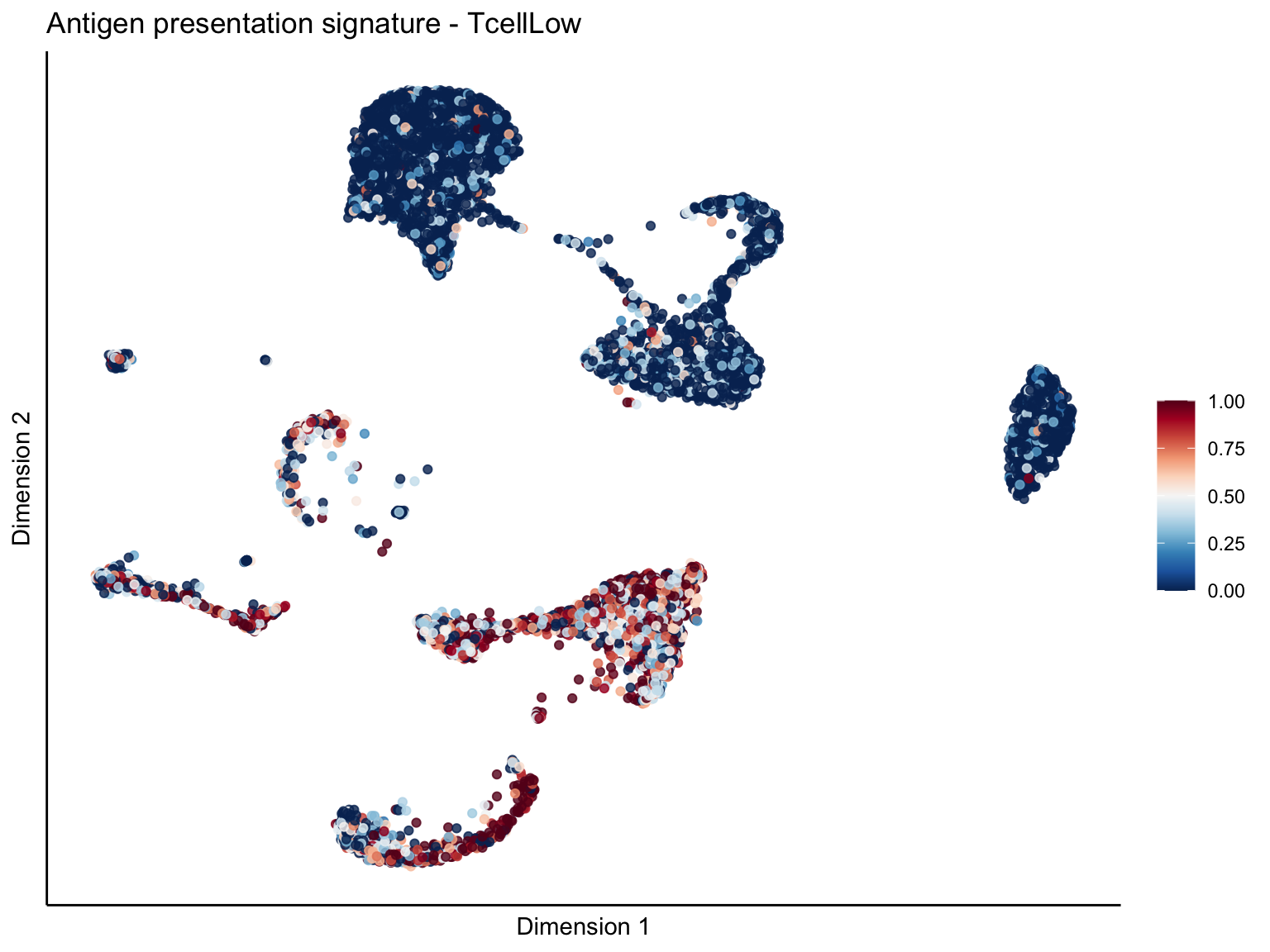
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]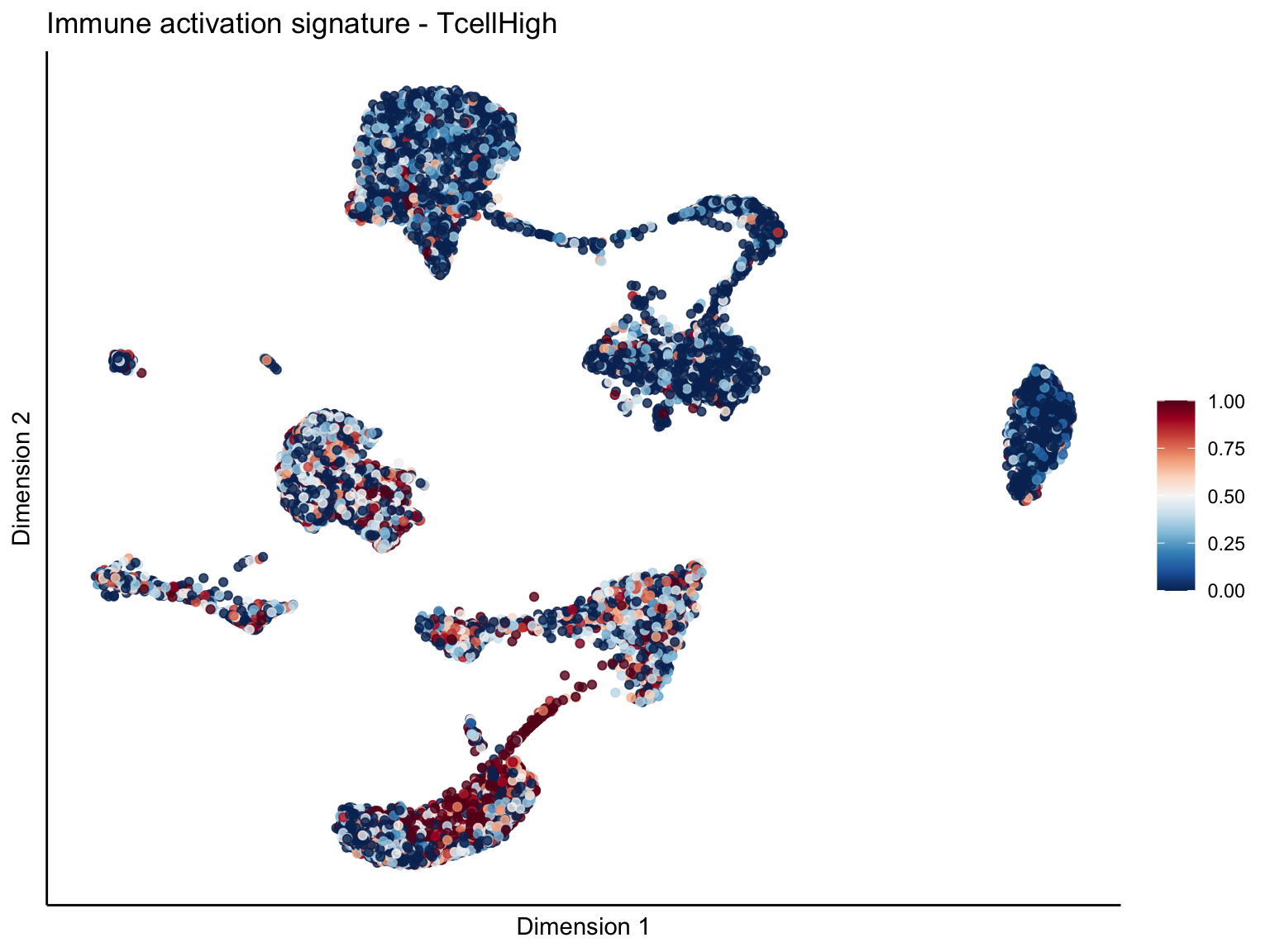
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]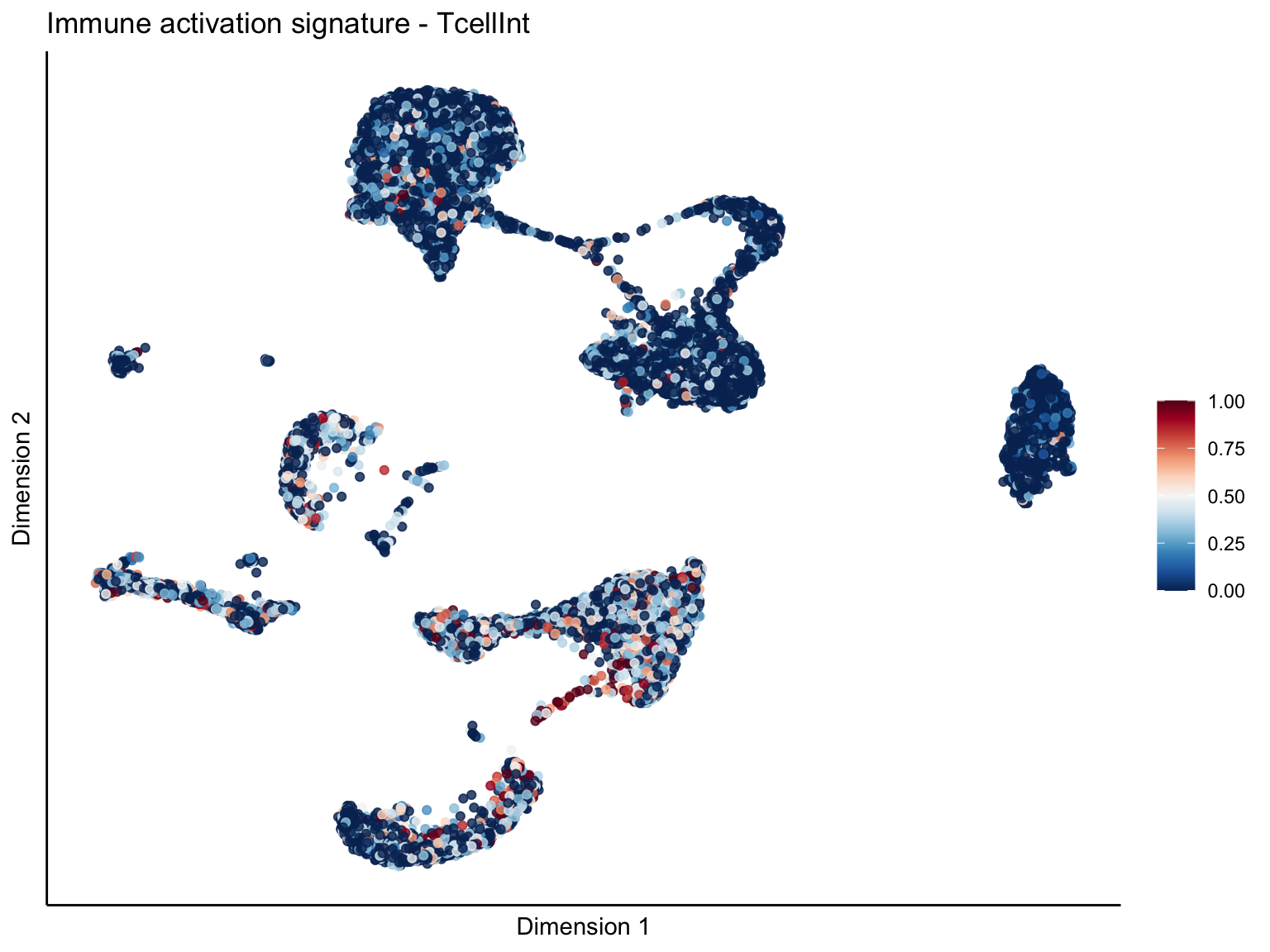
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]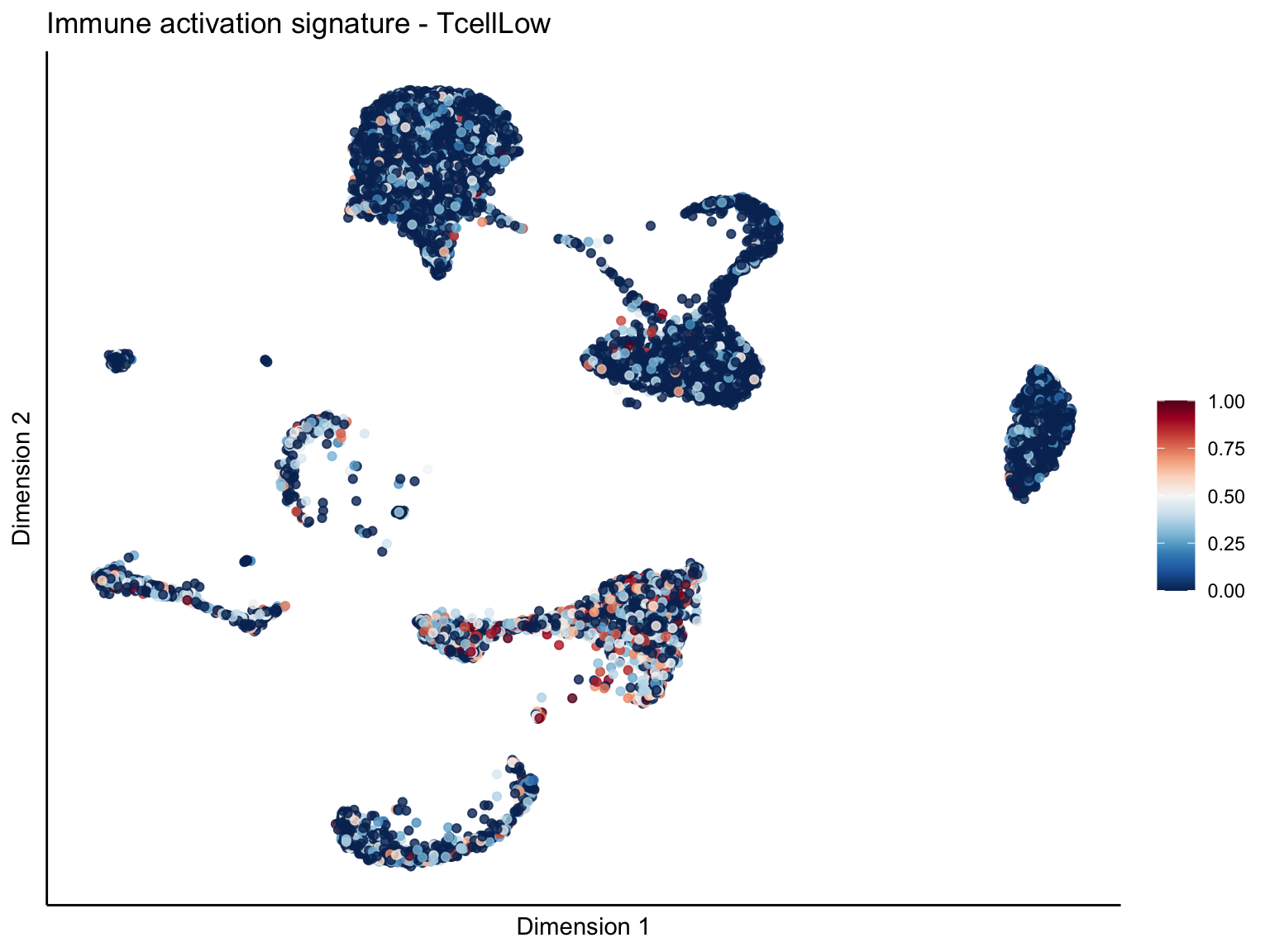
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]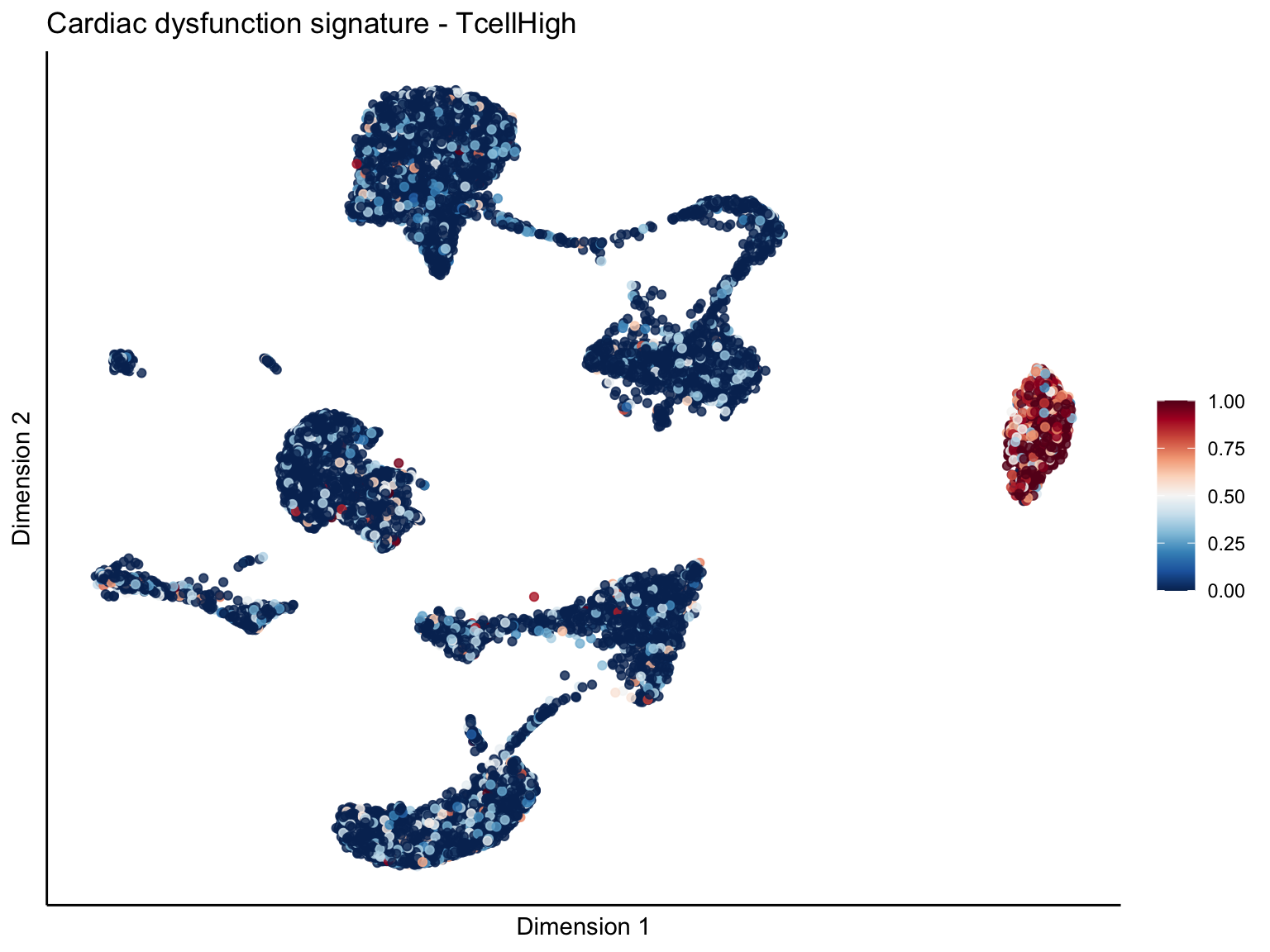
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]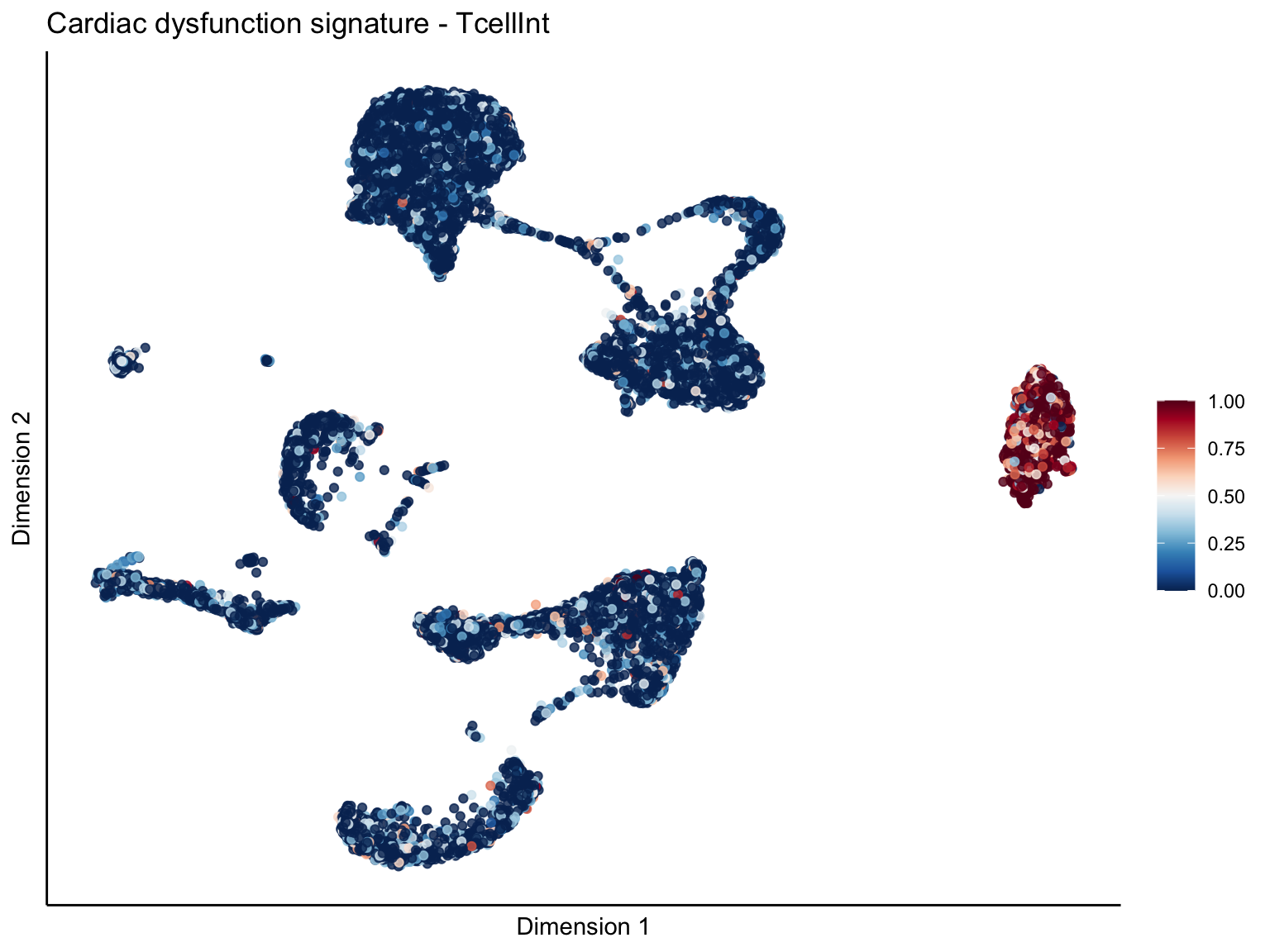
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]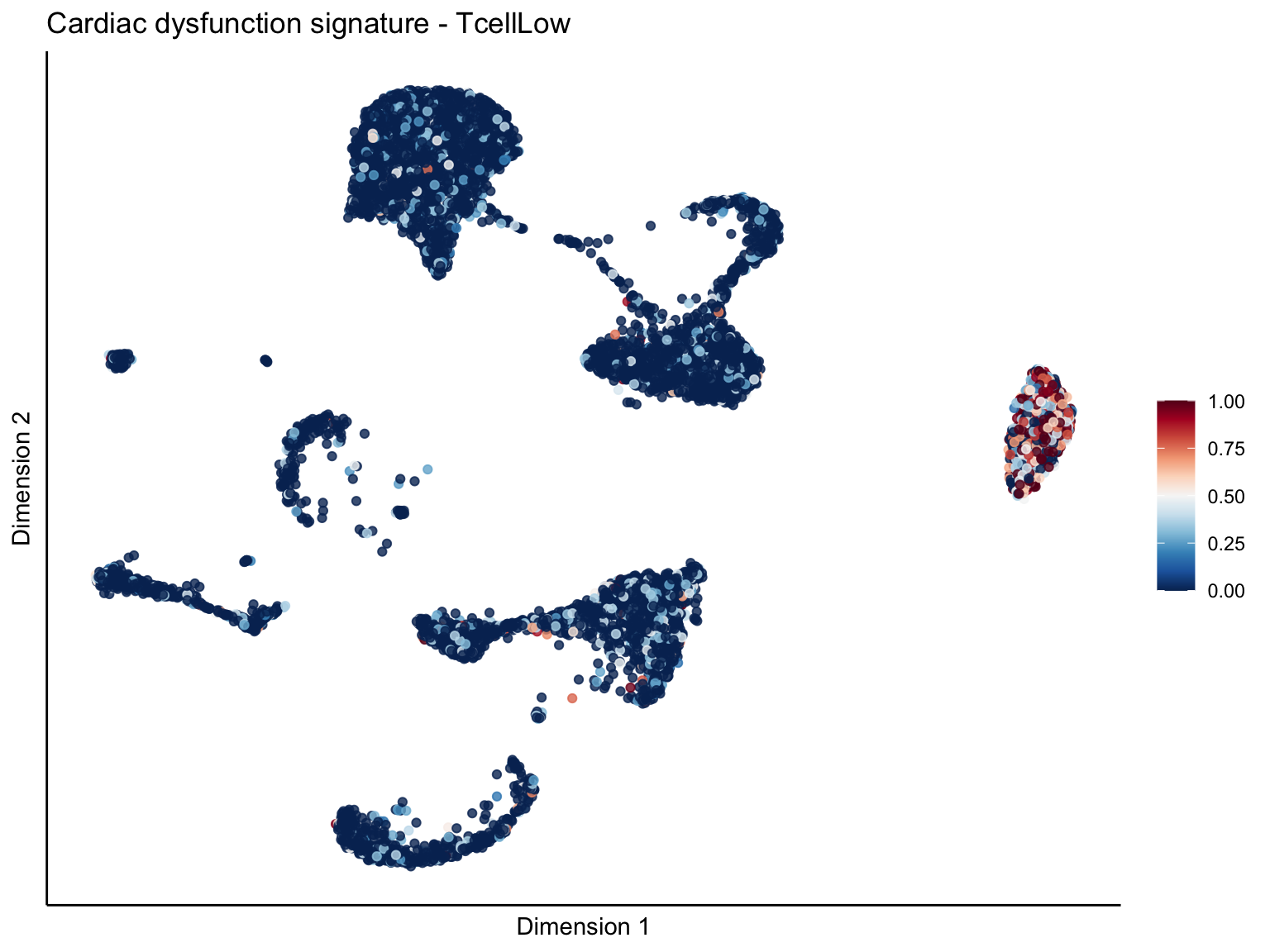
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]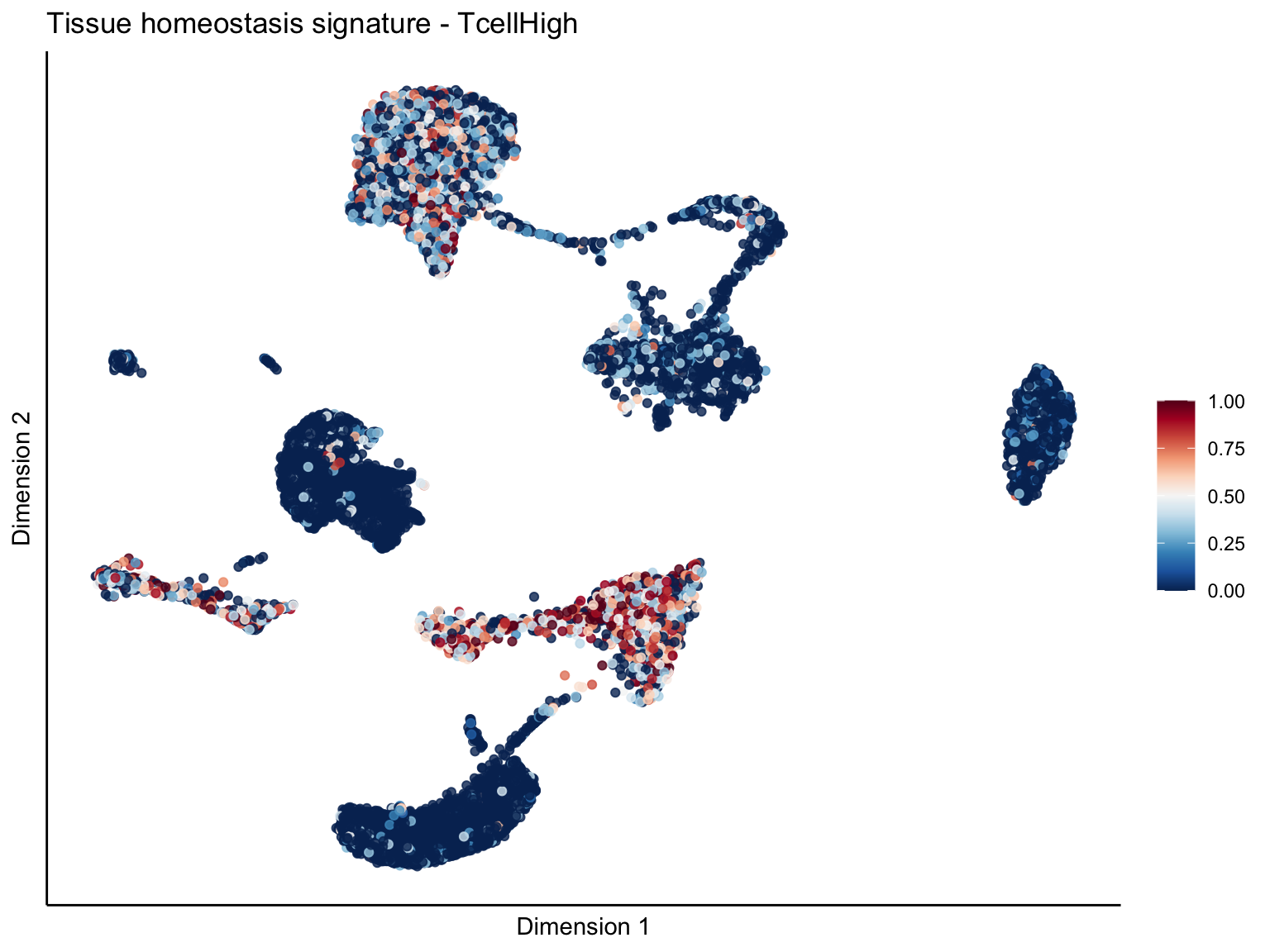
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]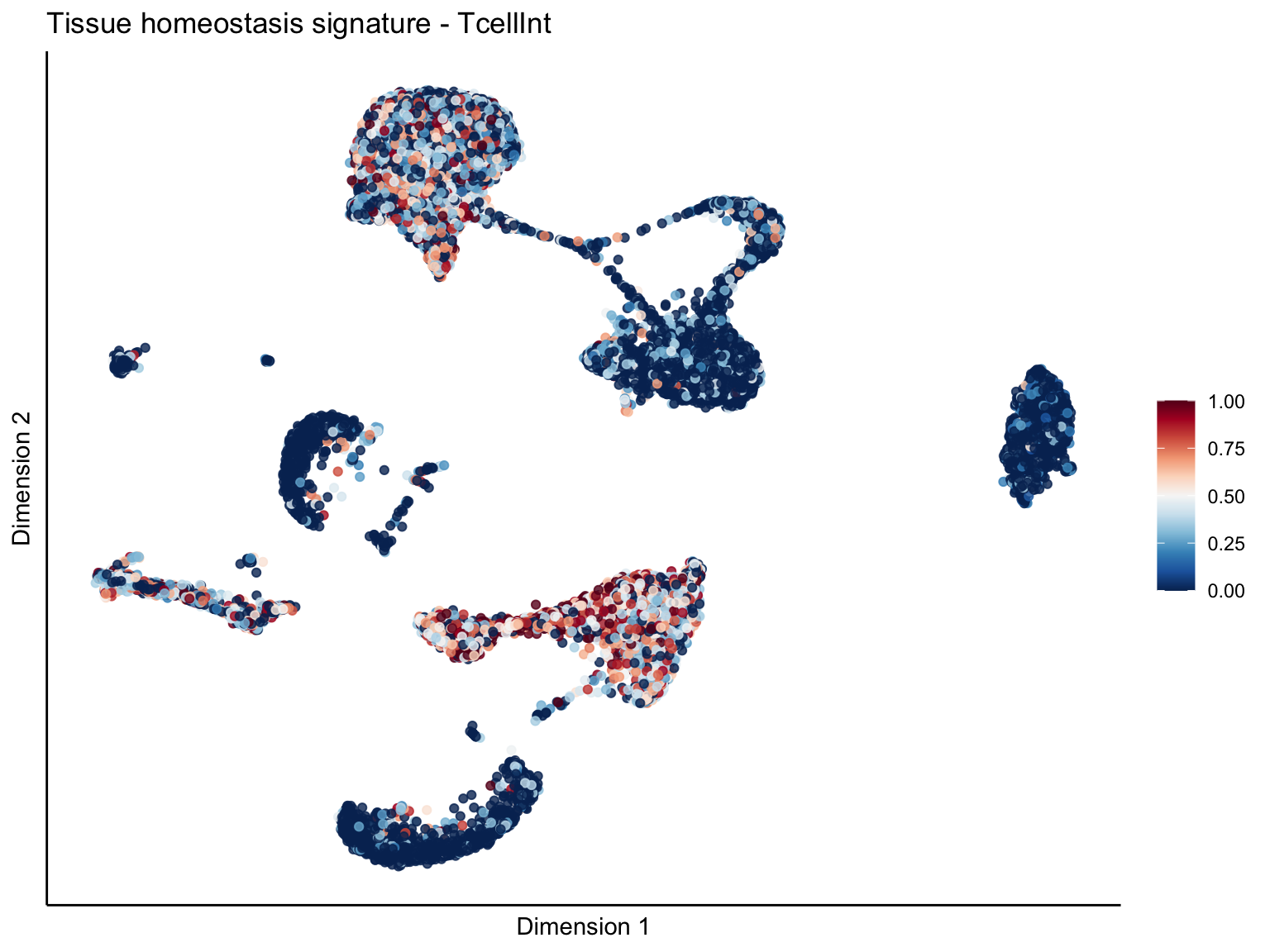
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]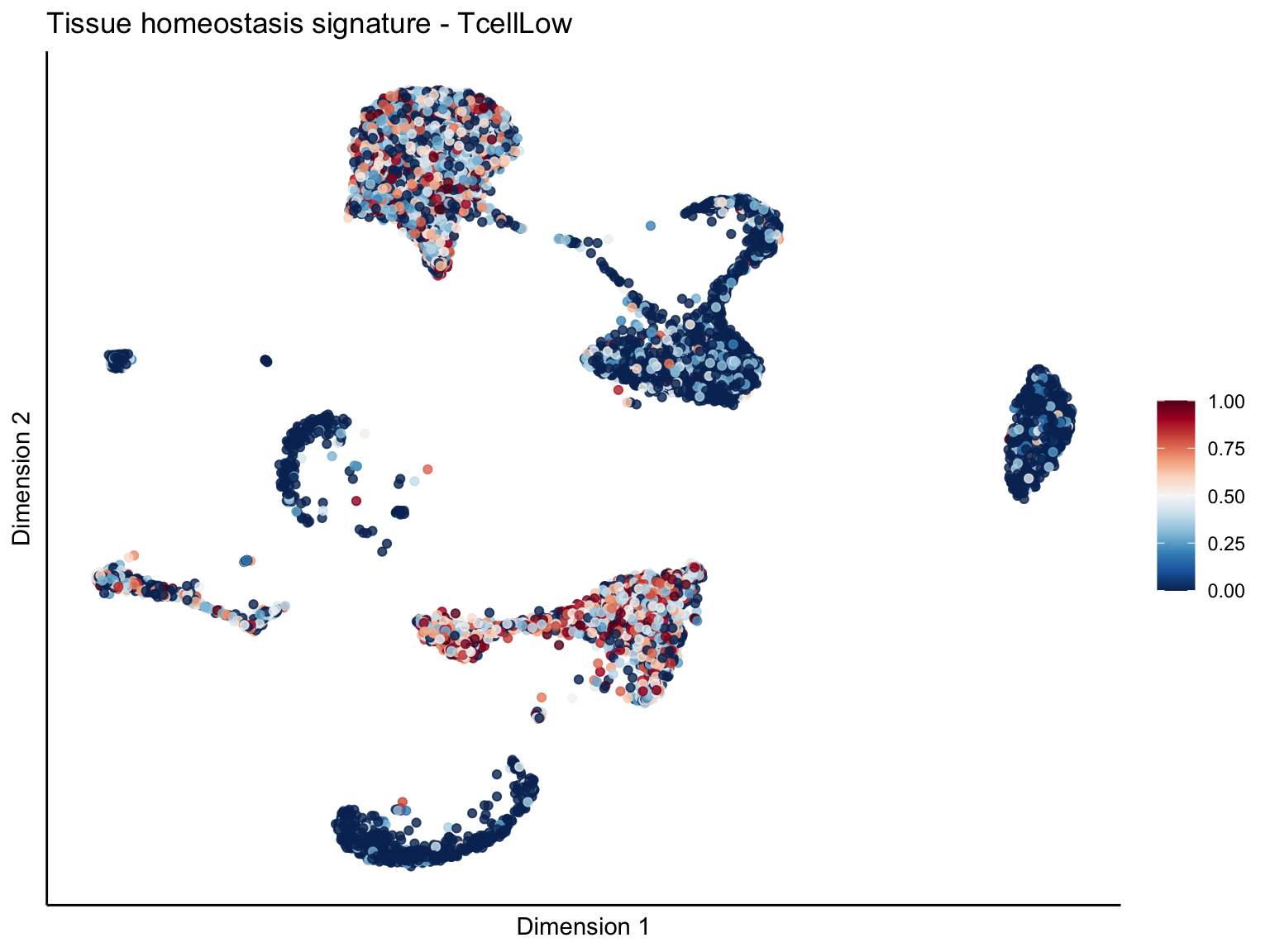
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
pal = viridis(100)
sc <- scale_colour_gradientn(colours = pal, limits=c(0, cutOff))
lapply(unique(signDat$signature), function(sign){
signGenes <- signDat %>% dplyr::filter(signature == sign)
sceSub <- sce[which(rownames(sce) %in% signGenes$geneID),]
cntMat <- rowSums(t(as.matrix(sceSub@assays@data$logcounts)))/nrow(signGenes)
sceSub$sign <- cntMat
sceSub$sign[which(sceSub$sign > cutOff)] <- cutOff
sceSub$sign[which(sceSub$sign < 0)] <- 0
lapply(treatGrps, function(treat){
sceSubT <- sceSub[, which(sceSub$TcellGrp == treat)]
p <- visGroup_adapt(sceSubT, 'sign', dim_red = 'UMAP') +
sc +
guides(colour = guide_colourbar(title = '')) +
ggtitle(paste0(sign, ' signature - ', treat)) +
theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank()) +
labs(x='Dimension 1', y='Dimension 2')
p
})
})[[1]]
[[1]][[1]]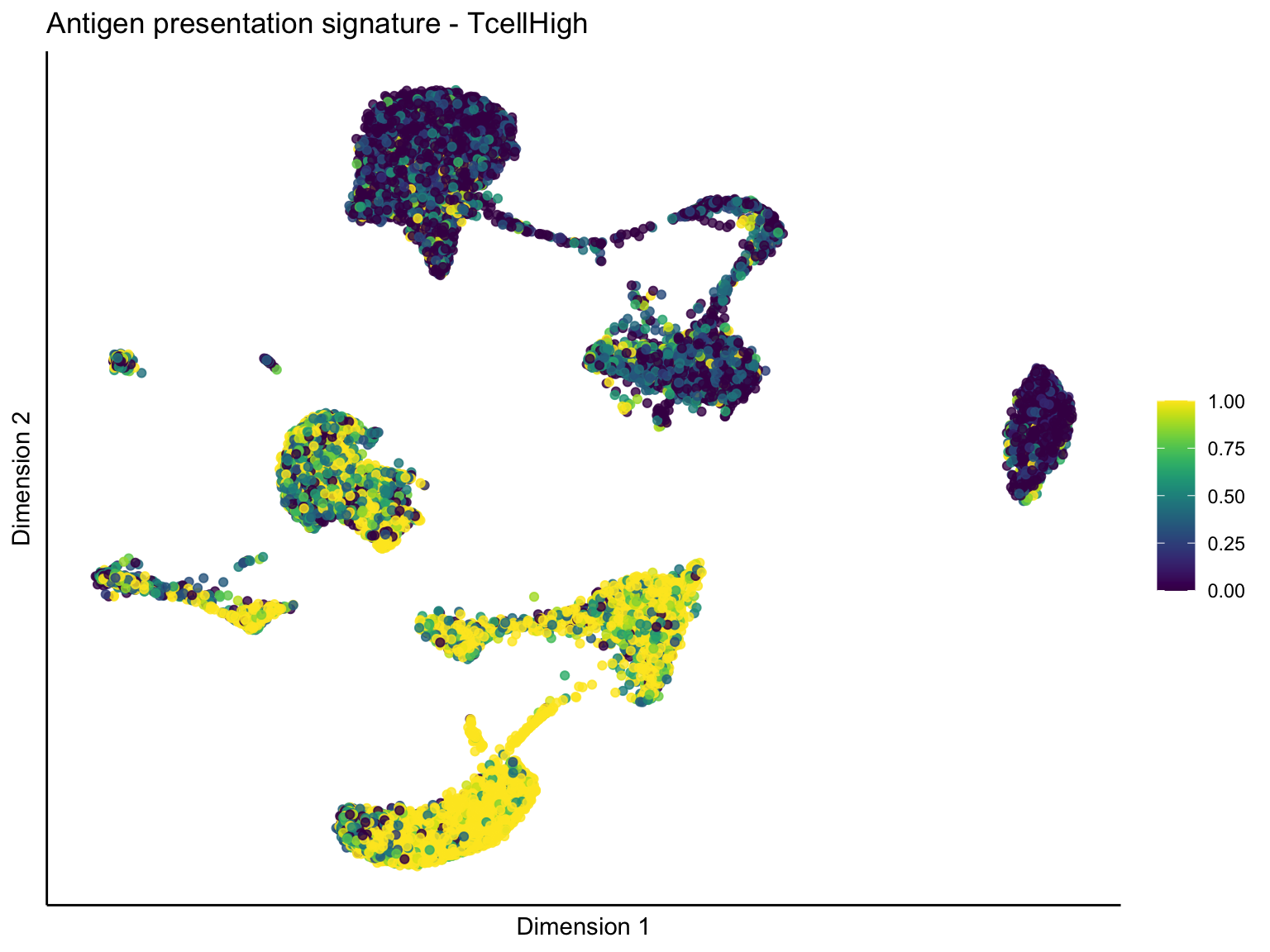
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[2]]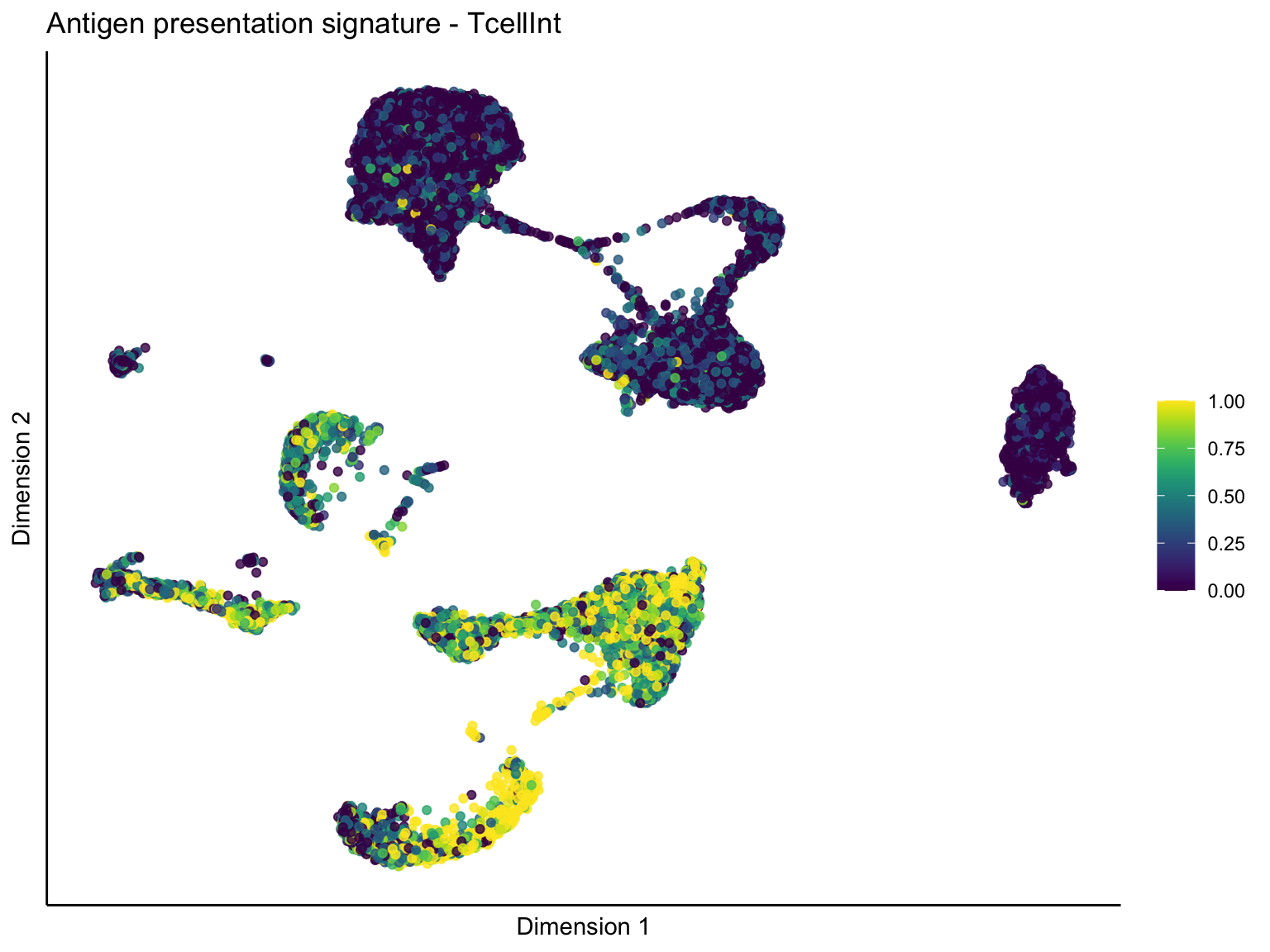
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[1]][[3]]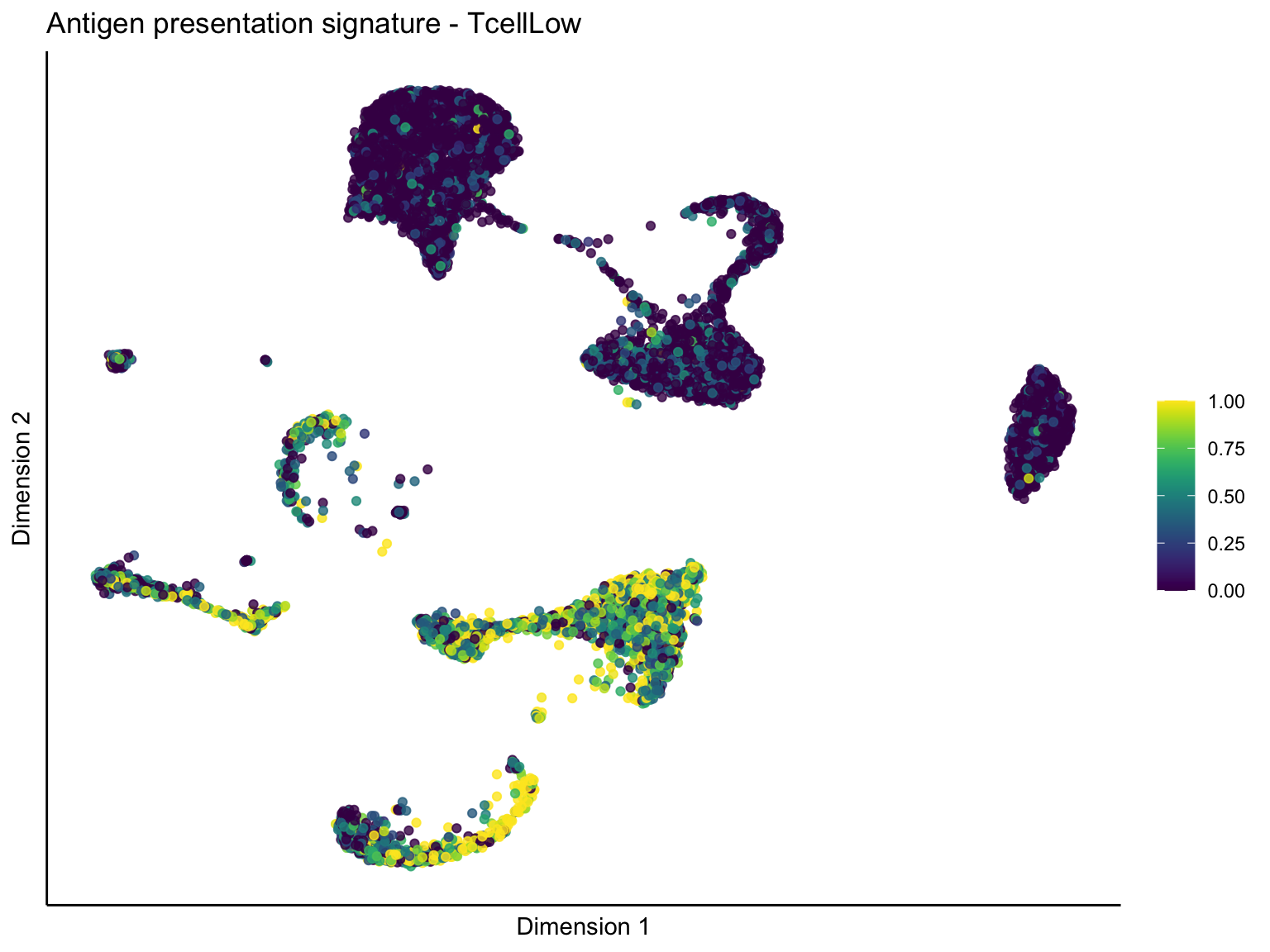
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]]
[[2]][[1]]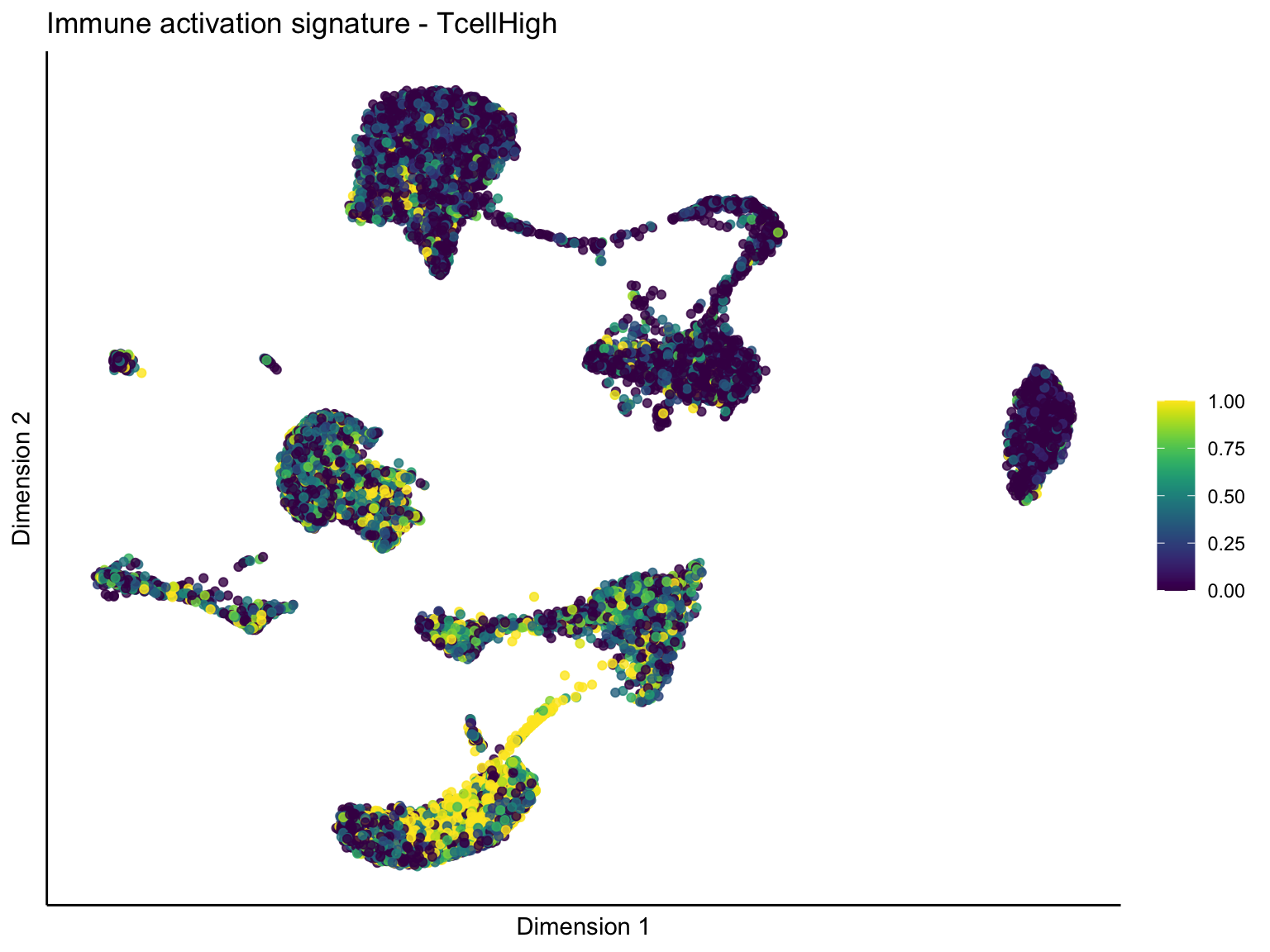
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[2]]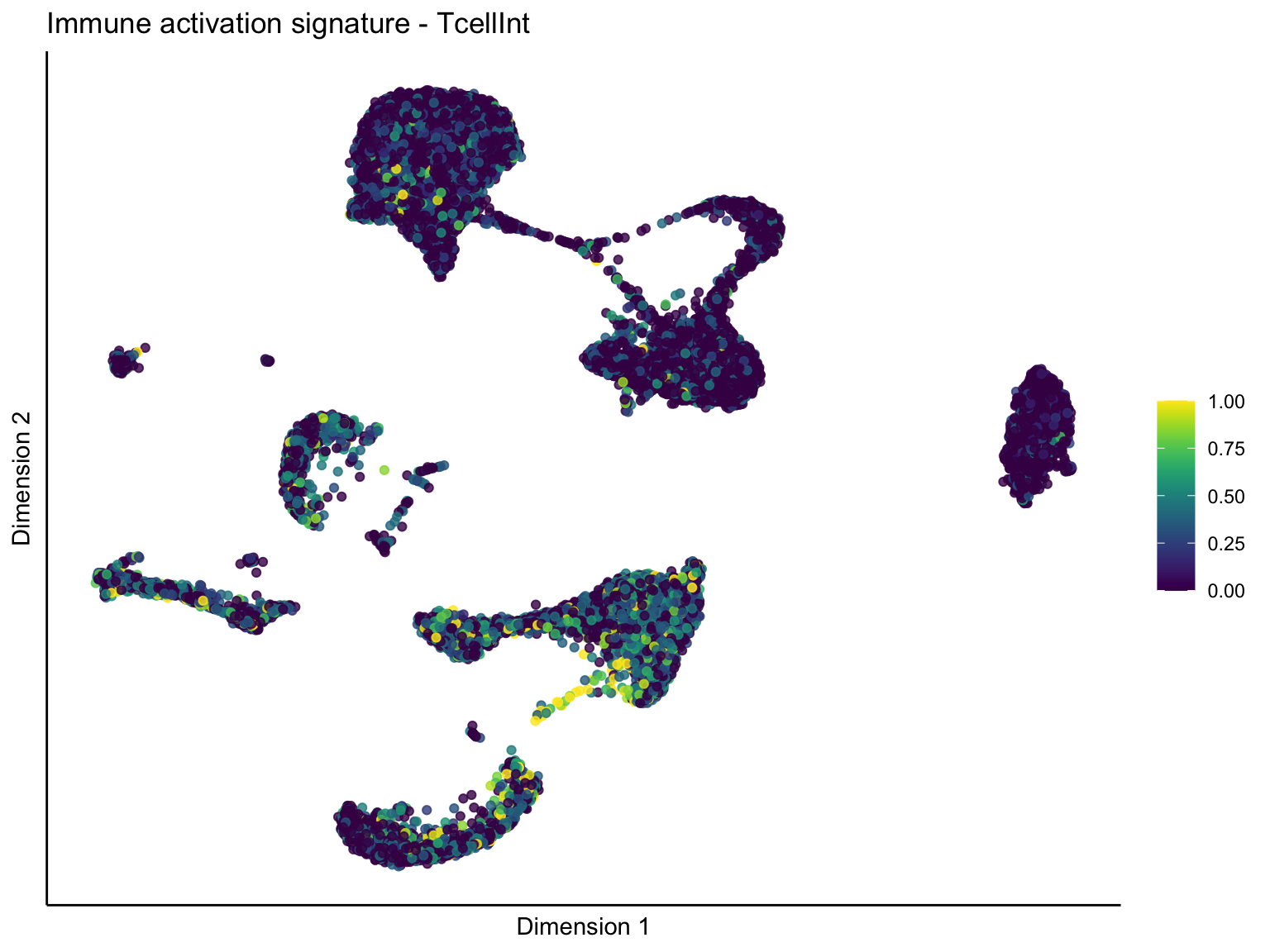
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[2]][[3]]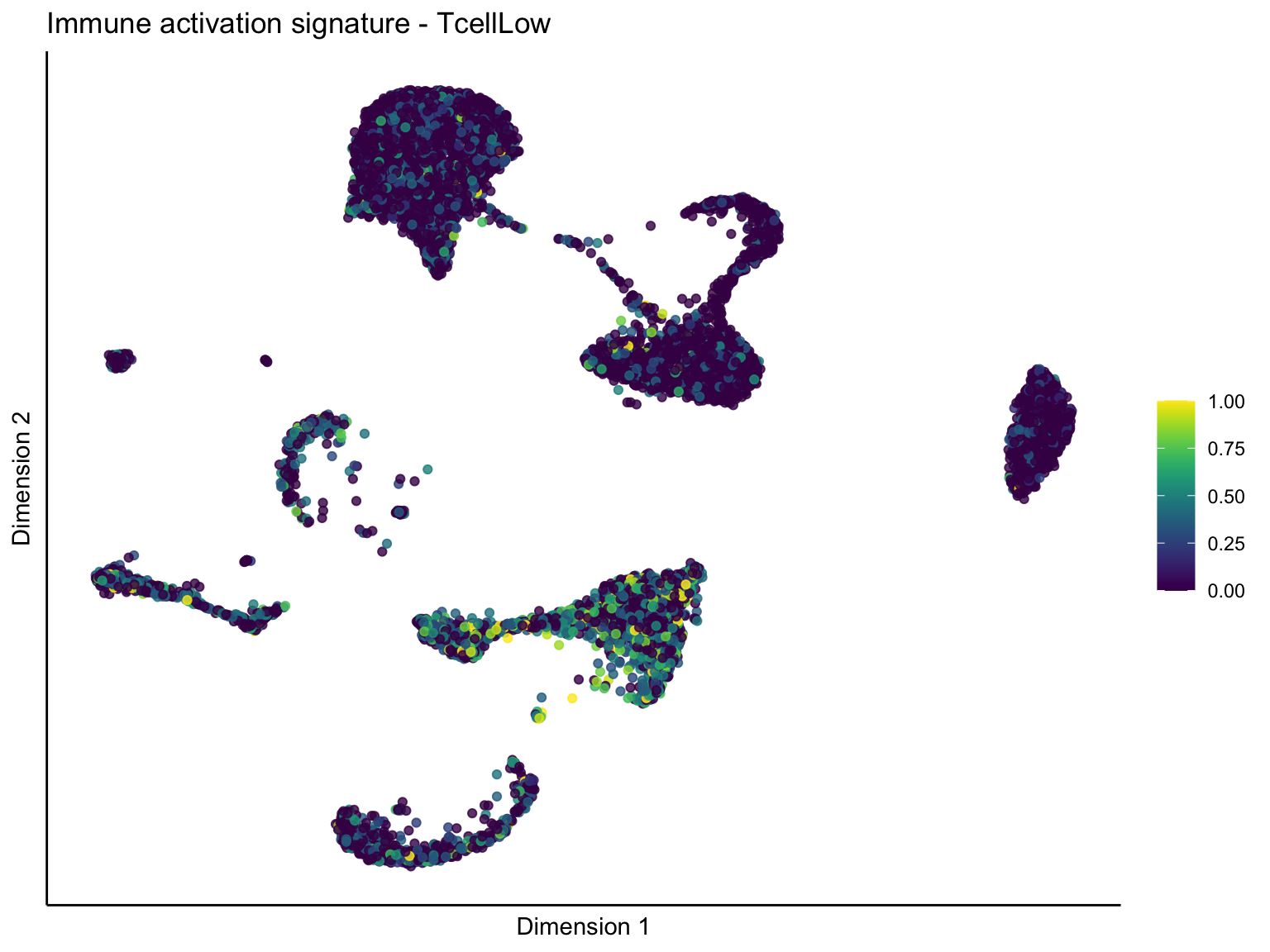
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]]
[[3]][[1]]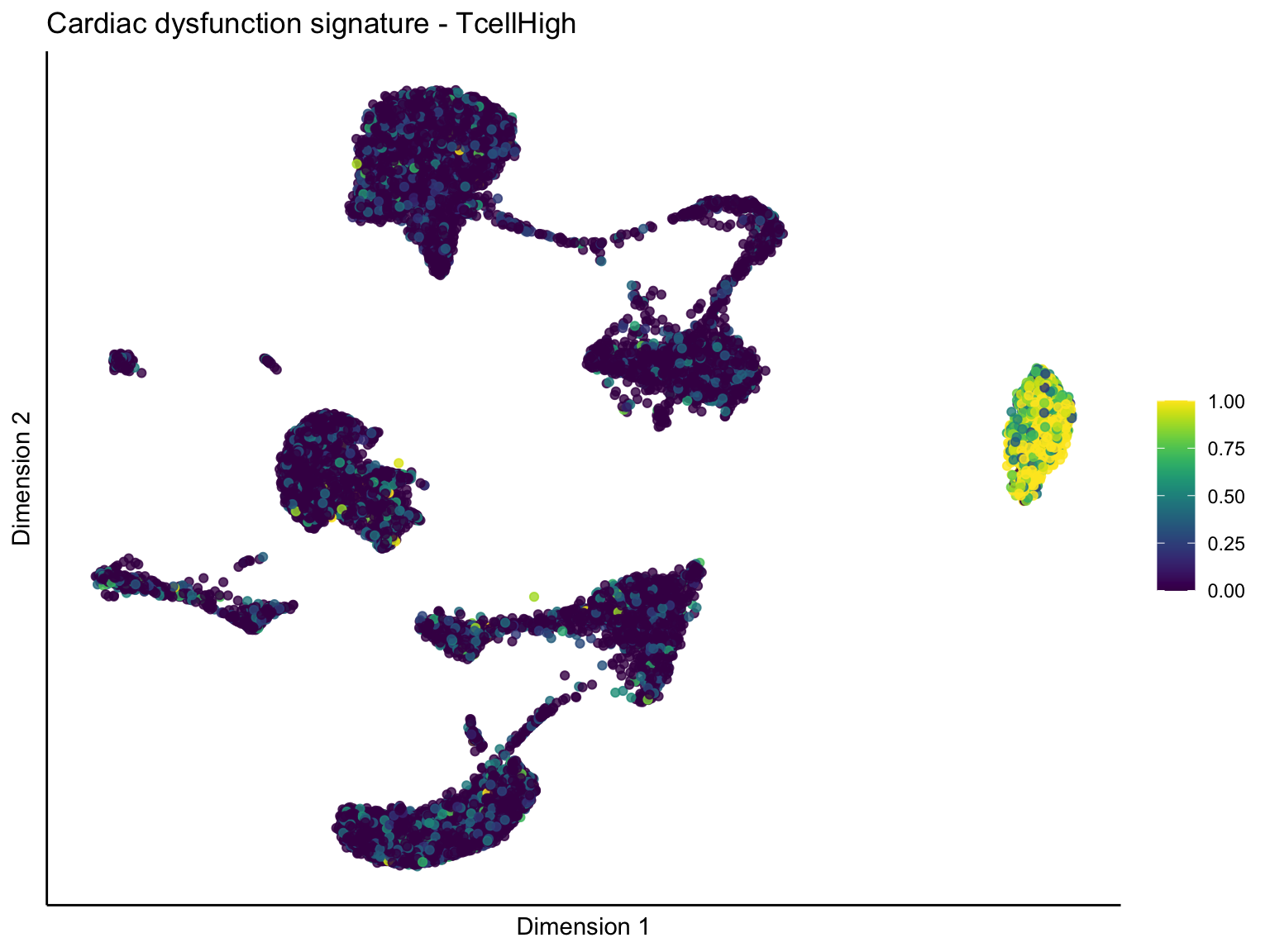
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[2]]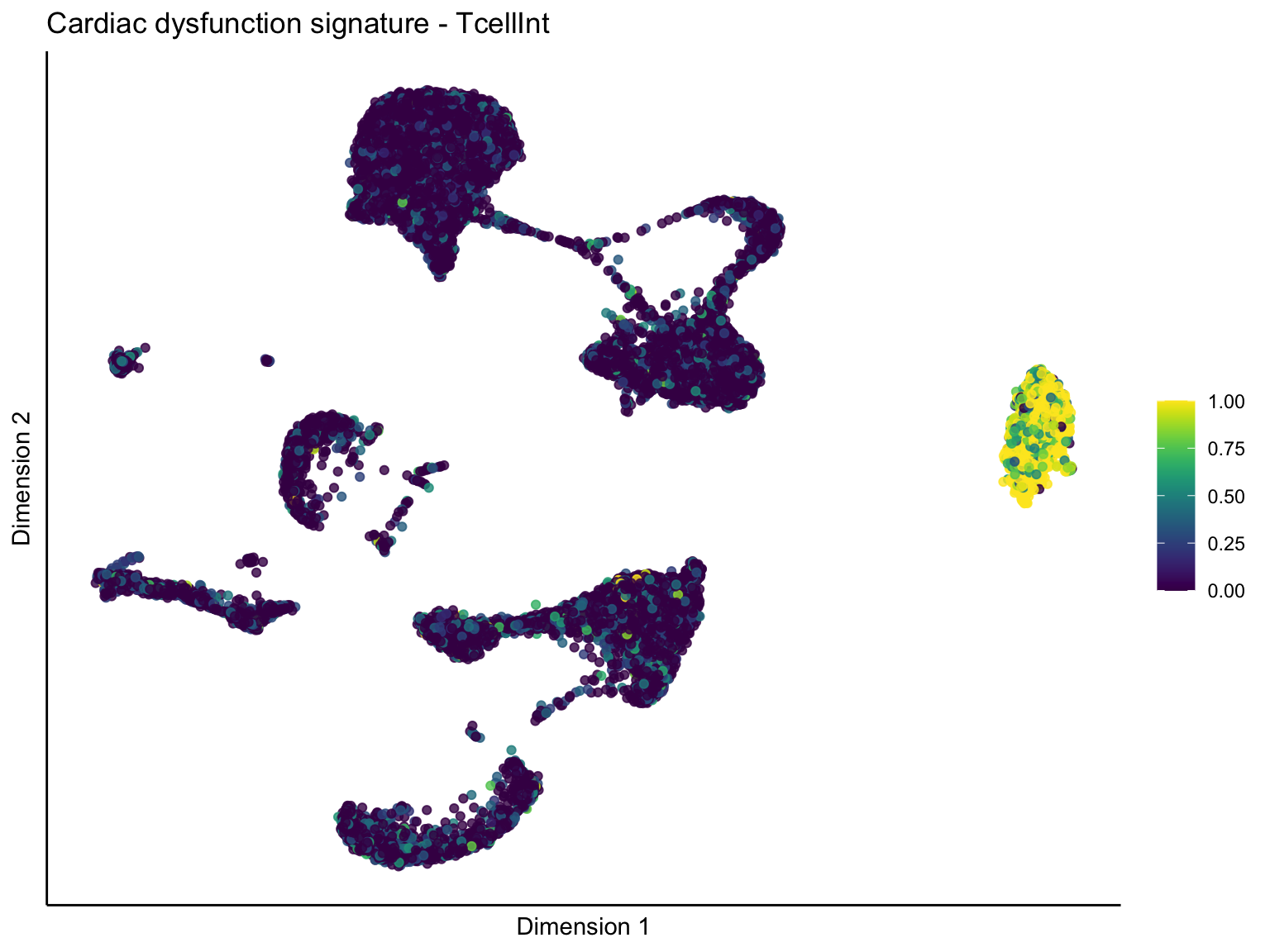
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[3]][[3]]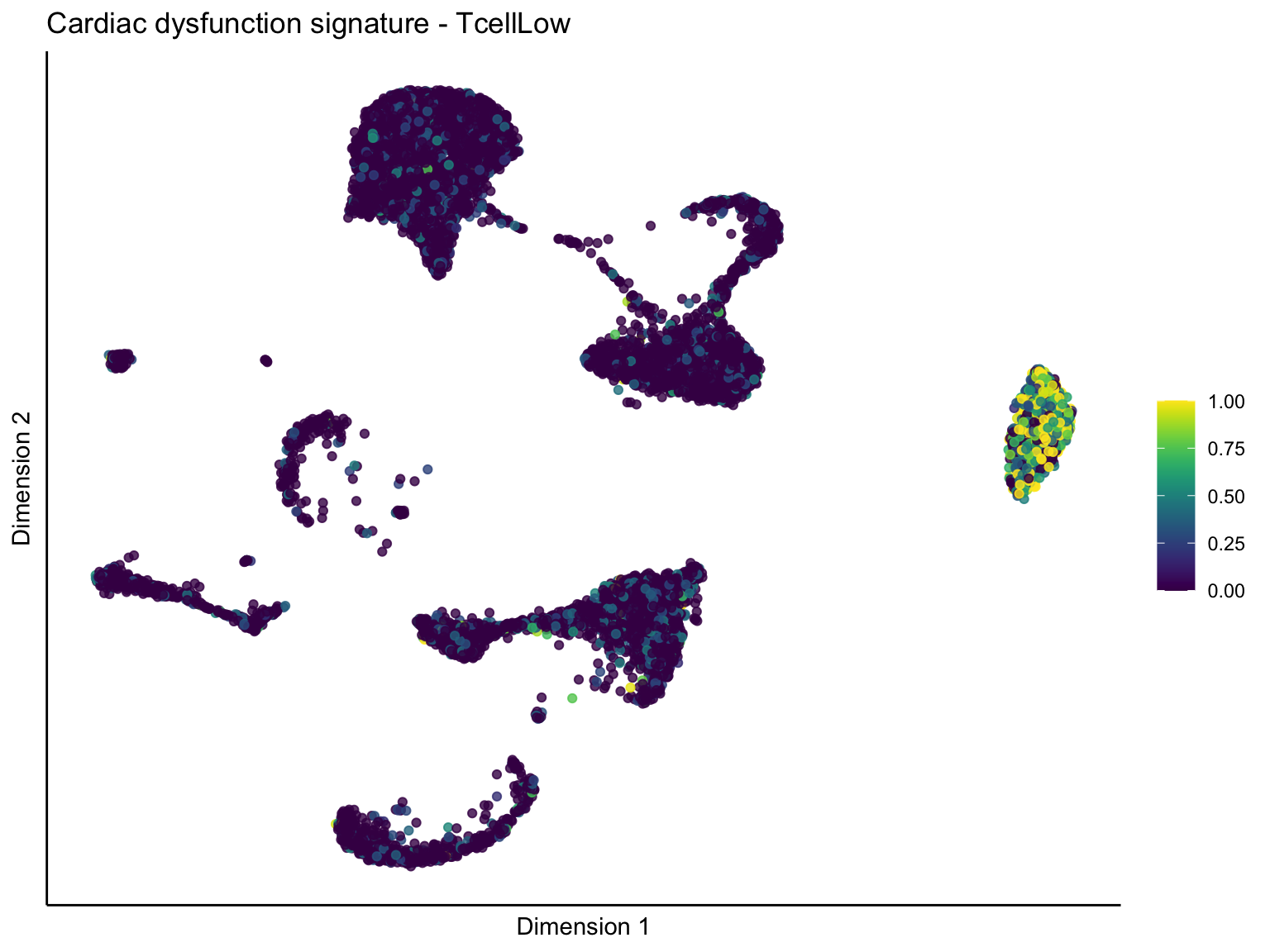
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]]
[[4]][[1]]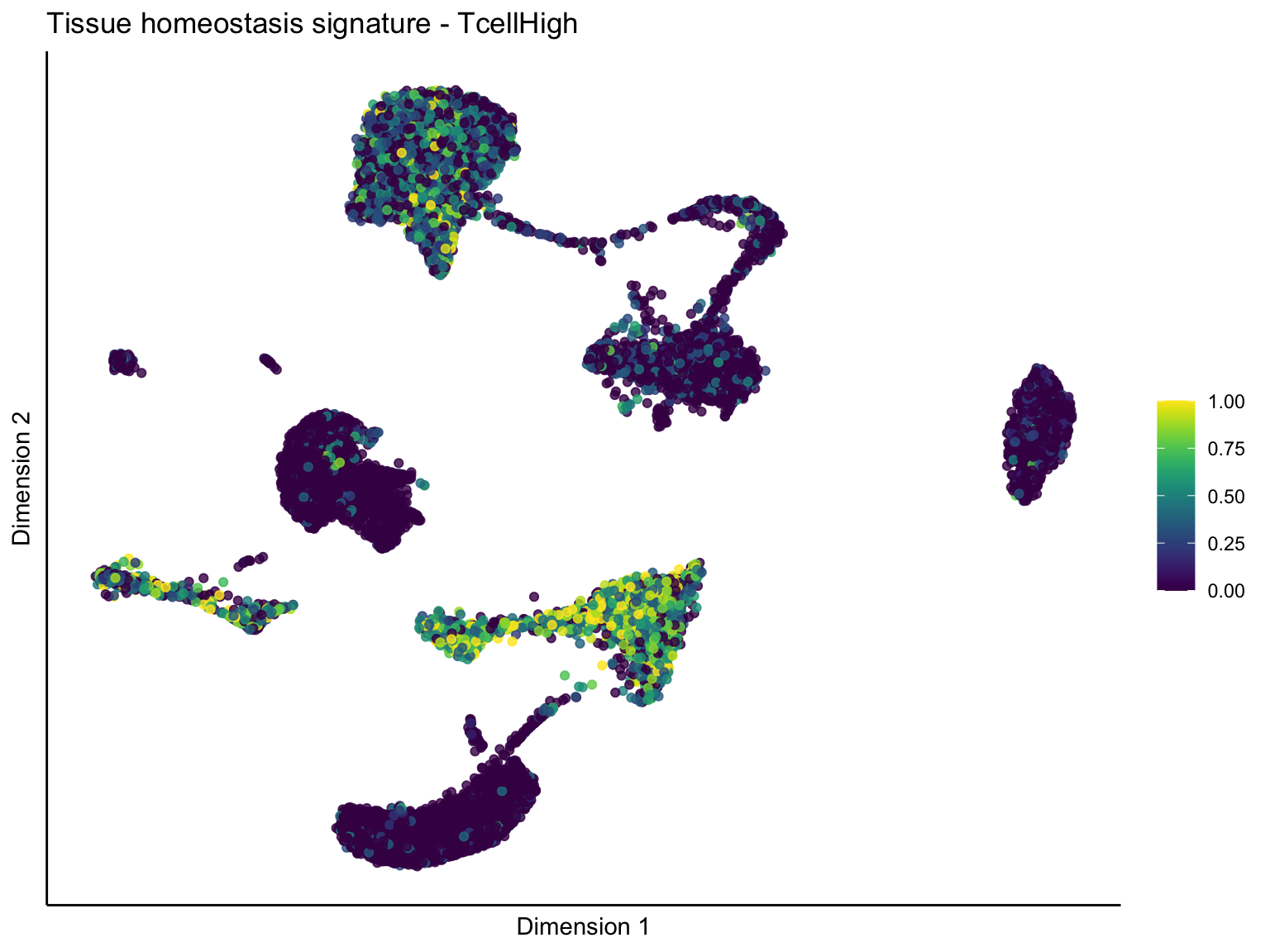
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[2]]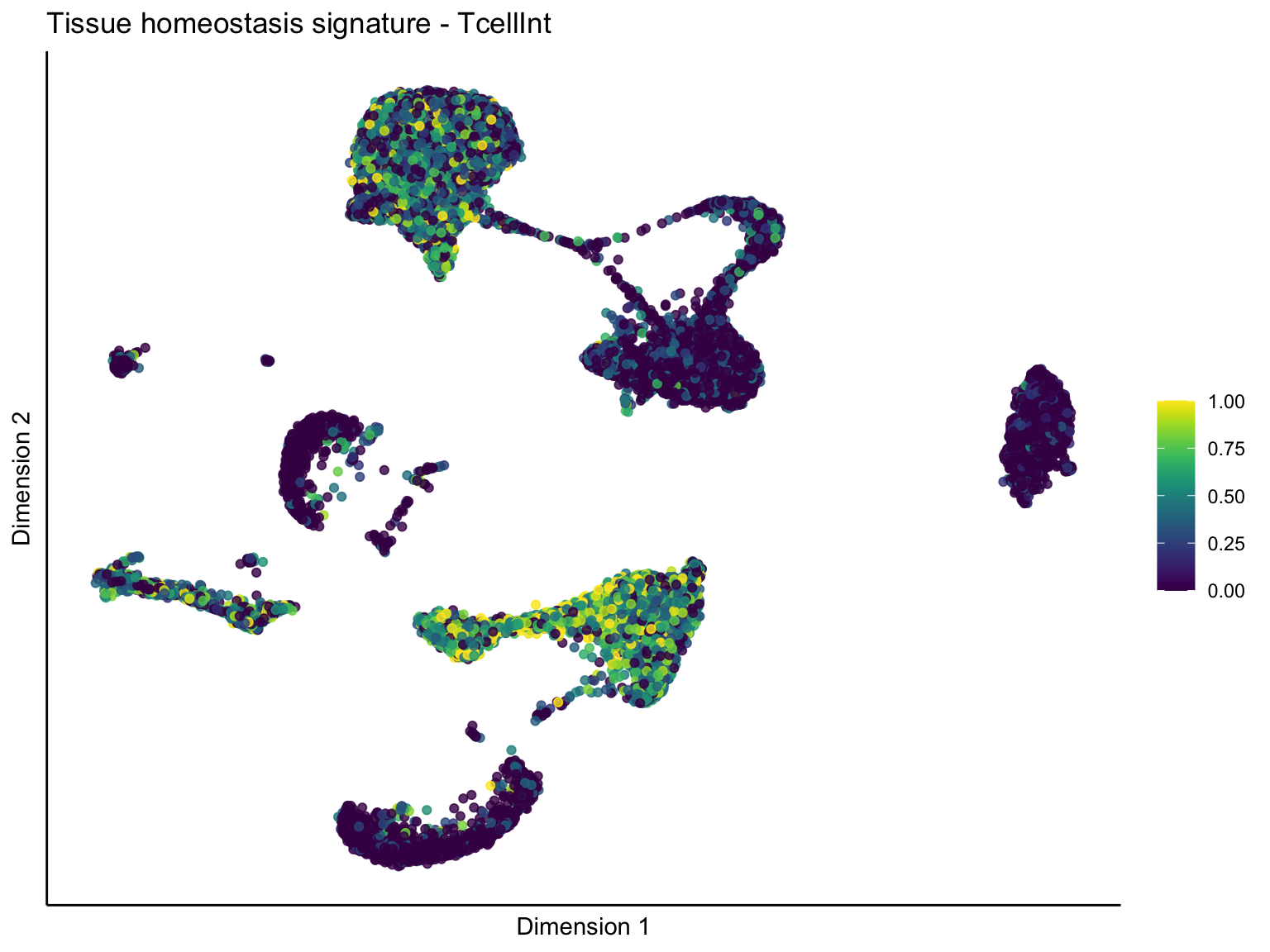
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
[[4]][[3]]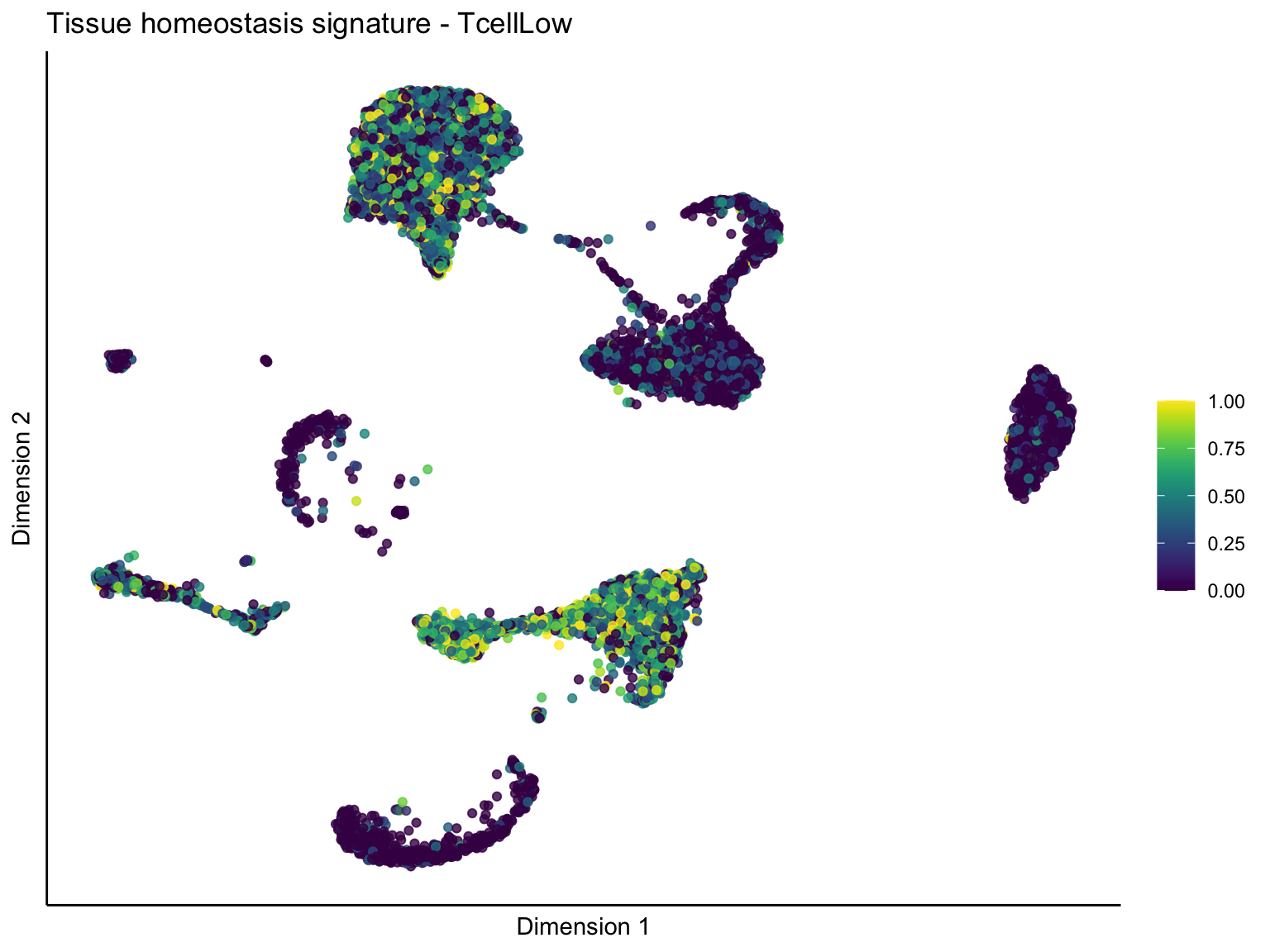
| Version | Author | Date |
|---|---|---|
| 66c2208 | mluetge | 2022-12-08 |
project signatures on heatmap
signDat <- read_delim(file = paste0(basedir,
"/data/GSEA/selGenesSignature2.txt"),
delim = "\t")
genes <- data.frame(geneID=rownames(seurat)) %>%
mutate(gene=gsub("^.*\\.", "", geneID))
signDat <- signDat %>% left_join(.,genes, by="gene")
## only CM, Fibroblasts, Tcells and Myeloids
selLabel <- c("Tcell","resMacrophage", "Fibroblast","infMacrophage",
"Cardiomyocyte")
seurat <- subset(seurat, label %in% selLabel)
seurat$label2 <- seurat$label
seurat$label2[which(seurat$label %in% c("resMacrophage",
"infMacrophage"))] <- "Macrophage"
seurat$label2_plus_grp <- paste0(seurat$label2, "_", seurat$TcellGrp)
table(seurat$label2_plus_grp)
Cardiomyocyte_TcellHigh Cardiomyocyte_TcellInt Cardiomyocyte_TcellLow
1023 1338 871
Fibroblast_TcellHigh Fibroblast_TcellInt Fibroblast_TcellLow
2894 5120 3162
Macrophage_TcellHigh Macrophage_TcellInt Macrophage_TcellLow
3530 1162 604
Tcell_TcellHigh Tcell_TcellInt Tcell_TcellLow
4369 768 231 seurat$label2_plus_grp <- as.factor(seurat$label2_plus_grp)
Idents(seurat) <- seurat$label2_plus_grp
gapVecCol <- seq(3, length(levels(seurat$label2_plus_grp)), by=3)
gapVecDat <- signDat %>% group_by(signature) %>% summarise(cnt=n())
gapVecRow <- cumsum(gapVecDat$cnt)
colLab2 <- c("#ba4e45", "#d4cc84", "#546f82", "#8d5639")
names(colLab2) <- c("Tcell","Macrophage","Fibroblast","Cardiomyocyte")
pOut <- avgHeatmap(seurat = seurat, selGenes = signDat,
colVecIdent = colLab2, colVecCond=colTgrp,
ordVec=levels(seurat),
gapVecR=gapVecRow, gapVecC=gapVecCol,cc=FALSE,
cr=F, condCol=T)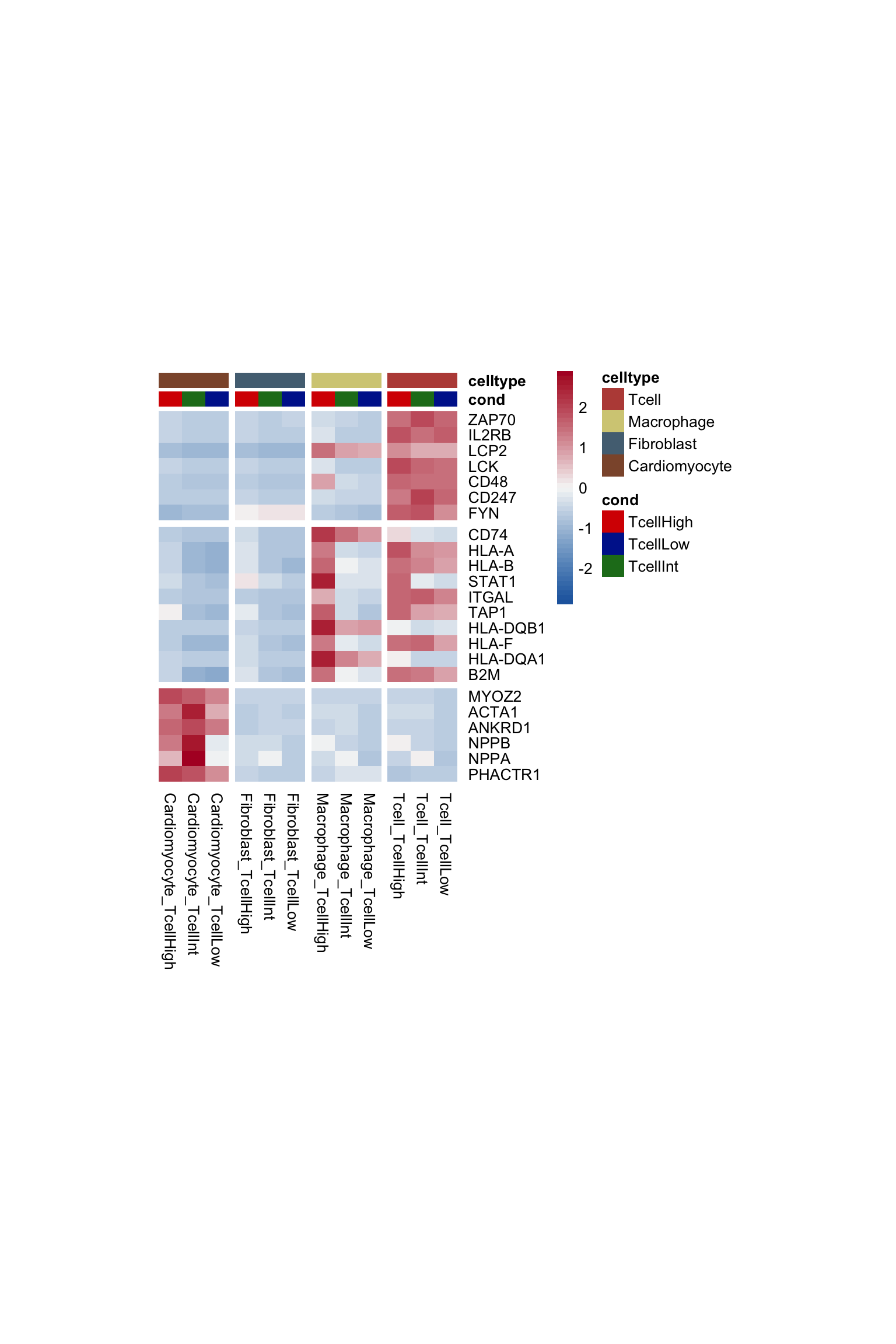
| Version | Author | Date |
|---|---|---|
| 307a2ca | mluetge | 2022-12-12 |
seurat <- subset(seurat, TcellGrp %in% c("TcellHigh", "TcellLow"))
gapVecCol <- seq(2, length(unique(seurat$label2_plus_grp)), by=2)
gapVecDat <- signDat %>% group_by(signature) %>% summarise(cnt=n())
gapVecRow <- cumsum(gapVecDat$cnt)
pOut <- avgHeatmap(seurat = seurat, selGenes = signDat,
colVecIdent = colLab2, colVecCond=colTgrp,
ordVec=levels(seurat),
gapVecR=gapVecRow, gapVecC=gapVecCol,cc=FALSE,
cr=F, condCol=T)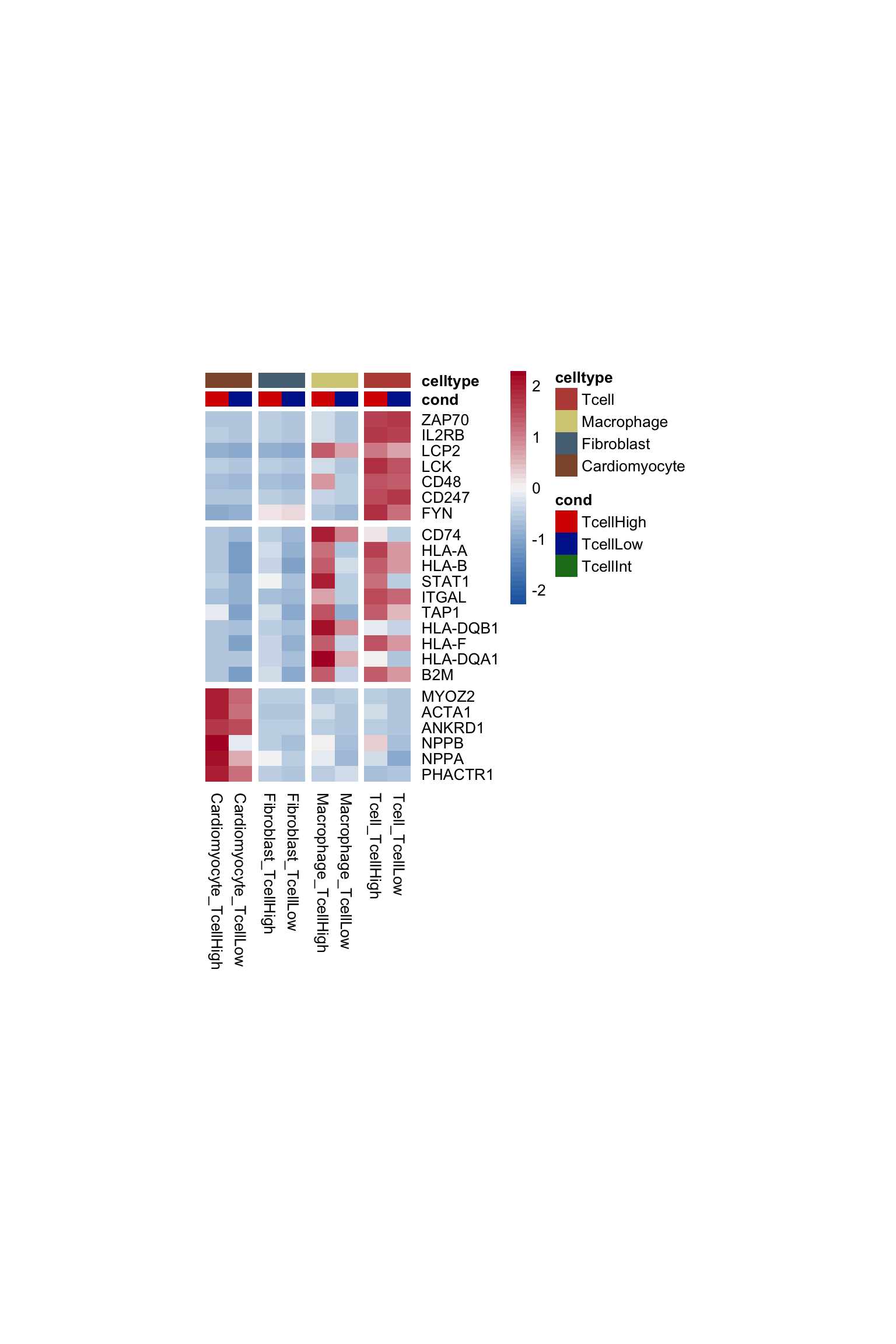
session info
sessionInfo()R version 4.2.1 (2022-06-23)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur ... 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] viridis_0.6.2 viridisLite_0.4.1
[3] RColorBrewer_1.1-3 ggpubr_0.5.0
[5] pheatmap_1.0.12 ggsci_2.9
[7] runSeurat3_0.1.0 here_1.0.1
[9] magrittr_2.0.3 SeuratObject_4.1.3
[11] Seurat_4.3.0 forcats_0.5.2
[13] stringr_1.5.0 dplyr_1.0.10
[15] purrr_0.3.5 readr_2.1.3
[17] tidyr_1.2.1 tibble_3.1.8
[19] ggplot2_3.4.0 tidyverse_1.3.2
[21] SingleCellExperiment_1.18.1 SummarizedExperiment_1.26.1
[23] Biobase_2.56.0 GenomicRanges_1.48.0
[25] GenomeInfoDb_1.32.4 IRanges_2.30.1
[27] S4Vectors_0.34.0 BiocGenerics_0.42.0
[29] MatrixGenerics_1.8.1 matrixStats_0.63.0
loaded via a namespace (and not attached):
[1] utf8_1.2.2 spatstat.explore_3.0-5 reticulate_1.26
[4] tidyselect_1.2.0 htmlwidgets_1.5.4 grid_4.2.1
[7] Rtsne_0.16 munsell_0.5.0 codetools_0.2-18
[10] ica_1.0-3 future_1.29.0 miniUI_0.1.1.1
[13] withr_2.5.0 spatstat.random_3.0-1 colorspace_2.0-3
[16] progressr_0.11.0 highr_0.9 knitr_1.41
[19] rstudioapi_0.14 ROCR_1.0-11 ggsignif_0.6.4
[22] tensor_1.5 listenv_0.8.0 labeling_0.4.2
[25] git2r_0.30.1 GenomeInfoDbData_1.2.8 polyclip_1.10-4
[28] bit64_4.0.5 farver_2.1.1 rprojroot_2.0.3
[31] parallelly_1.32.1 vctrs_0.5.1 generics_0.1.3
[34] xfun_0.35 timechange_0.1.1 R6_2.5.1
[37] bitops_1.0-7 spatstat.utils_3.0-1 cachem_1.0.6
[40] DelayedArray_0.22.0 assertthat_0.2.1 vroom_1.6.0
[43] promises_1.2.0.1 scales_1.2.1 googlesheets4_1.0.1
[46] gtable_0.3.1 globals_0.16.2 goftest_1.2-3
[49] workflowr_1.7.0 rlang_1.0.6 splines_4.2.1
[52] rstatix_0.7.1 lazyeval_0.2.2 gargle_1.2.1
[55] spatstat.geom_3.0-3 broom_1.0.1 yaml_2.3.6
[58] reshape2_1.4.4 abind_1.4-5 modelr_0.1.10
[61] backports_1.4.1 httpuv_1.6.6 tools_4.2.1
[64] ellipsis_0.3.2 jquerylib_0.1.4 ggridges_0.5.4
[67] Rcpp_1.0.9 plyr_1.8.8 zlibbioc_1.42.0
[70] RCurl_1.98-1.9 deldir_1.0-6 pbapply_1.6-0
[73] cowplot_1.1.1 zoo_1.8-11 haven_2.5.1
[76] ggrepel_0.9.2 cluster_2.1.4 fs_1.5.2
[79] data.table_1.14.6 scattermore_0.8 lmtest_0.9-40
[82] reprex_2.0.2 RANN_2.6.1 googledrive_2.0.0
[85] whisker_0.4 fitdistrplus_1.1-8 hms_1.1.2
[88] patchwork_1.1.2 mime_0.12 evaluate_0.18
[91] xtable_1.8-4 readxl_1.4.1 gridExtra_2.3
[94] compiler_4.2.1 KernSmooth_2.23-20 crayon_1.5.2
[97] htmltools_0.5.3 later_1.3.0 tzdb_0.3.0
[100] lubridate_1.9.0 DBI_1.1.3 dbplyr_2.2.1
[103] MASS_7.3-58.1 Matrix_1.5-3 car_3.1-1
[106] cli_3.4.1 parallel_4.2.1 igraph_1.3.5
[109] pkgconfig_2.0.3 sp_1.5-1 plotly_4.10.1
[112] spatstat.sparse_3.0-0 xml2_1.3.3 bslib_0.4.1
[115] XVector_0.36.0 rvest_1.0.3 digest_0.6.30
[118] sctransform_0.3.5 RcppAnnoy_0.0.20 spatstat.data_3.0-0
[121] rmarkdown_2.18 cellranger_1.1.0 leiden_0.4.3
[124] uwot_0.1.14 shiny_1.7.3 lifecycle_1.0.3
[127] nlme_3.1-160 jsonlite_1.8.3 carData_3.0-5
[130] fansi_1.0.3 pillar_1.8.1 lattice_0.20-45
[133] fastmap_1.1.0 httr_1.4.4 survival_3.4-0
[136] glue_1.6.2 png_0.1-8 bit_4.0.5
[139] stringi_1.7.8 sass_0.4.4 irlba_2.3.5.1
[142] future.apply_1.10.0 date()[1] "Mon Dec 12 17:18:39 2022"
sessionInfo()R version 4.2.1 (2022-06-23)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur ... 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] viridis_0.6.2 viridisLite_0.4.1
[3] RColorBrewer_1.1-3 ggpubr_0.5.0
[5] pheatmap_1.0.12 ggsci_2.9
[7] runSeurat3_0.1.0 here_1.0.1
[9] magrittr_2.0.3 SeuratObject_4.1.3
[11] Seurat_4.3.0 forcats_0.5.2
[13] stringr_1.5.0 dplyr_1.0.10
[15] purrr_0.3.5 readr_2.1.3
[17] tidyr_1.2.1 tibble_3.1.8
[19] ggplot2_3.4.0 tidyverse_1.3.2
[21] SingleCellExperiment_1.18.1 SummarizedExperiment_1.26.1
[23] Biobase_2.56.0 GenomicRanges_1.48.0
[25] GenomeInfoDb_1.32.4 IRanges_2.30.1
[27] S4Vectors_0.34.0 BiocGenerics_0.42.0
[29] MatrixGenerics_1.8.1 matrixStats_0.63.0
loaded via a namespace (and not attached):
[1] utf8_1.2.2 spatstat.explore_3.0-5 reticulate_1.26
[4] tidyselect_1.2.0 htmlwidgets_1.5.4 grid_4.2.1
[7] Rtsne_0.16 munsell_0.5.0 codetools_0.2-18
[10] ica_1.0-3 future_1.29.0 miniUI_0.1.1.1
[13] withr_2.5.0 spatstat.random_3.0-1 colorspace_2.0-3
[16] progressr_0.11.0 highr_0.9 knitr_1.41
[19] rstudioapi_0.14 ROCR_1.0-11 ggsignif_0.6.4
[22] tensor_1.5 listenv_0.8.0 labeling_0.4.2
[25] git2r_0.30.1 GenomeInfoDbData_1.2.8 polyclip_1.10-4
[28] bit64_4.0.5 farver_2.1.1 rprojroot_2.0.3
[31] parallelly_1.32.1 vctrs_0.5.1 generics_0.1.3
[34] xfun_0.35 timechange_0.1.1 R6_2.5.1
[37] bitops_1.0-7 spatstat.utils_3.0-1 cachem_1.0.6
[40] DelayedArray_0.22.0 assertthat_0.2.1 vroom_1.6.0
[43] promises_1.2.0.1 scales_1.2.1 googlesheets4_1.0.1
[46] gtable_0.3.1 globals_0.16.2 goftest_1.2-3
[49] workflowr_1.7.0 rlang_1.0.6 splines_4.2.1
[52] rstatix_0.7.1 lazyeval_0.2.2 gargle_1.2.1
[55] spatstat.geom_3.0-3 broom_1.0.1 yaml_2.3.6
[58] reshape2_1.4.4 abind_1.4-5 modelr_0.1.10
[61] backports_1.4.1 httpuv_1.6.6 tools_4.2.1
[64] ellipsis_0.3.2 jquerylib_0.1.4 ggridges_0.5.4
[67] Rcpp_1.0.9 plyr_1.8.8 zlibbioc_1.42.0
[70] RCurl_1.98-1.9 deldir_1.0-6 pbapply_1.6-0
[73] cowplot_1.1.1 zoo_1.8-11 haven_2.5.1
[76] ggrepel_0.9.2 cluster_2.1.4 fs_1.5.2
[79] data.table_1.14.6 scattermore_0.8 lmtest_0.9-40
[82] reprex_2.0.2 RANN_2.6.1 googledrive_2.0.0
[85] whisker_0.4 fitdistrplus_1.1-8 hms_1.1.2
[88] patchwork_1.1.2 mime_0.12 evaluate_0.18
[91] xtable_1.8-4 readxl_1.4.1 gridExtra_2.3
[94] compiler_4.2.1 KernSmooth_2.23-20 crayon_1.5.2
[97] htmltools_0.5.3 later_1.3.0 tzdb_0.3.0
[100] lubridate_1.9.0 DBI_1.1.3 dbplyr_2.2.1
[103] MASS_7.3-58.1 Matrix_1.5-3 car_3.1-1
[106] cli_3.4.1 parallel_4.2.1 igraph_1.3.5
[109] pkgconfig_2.0.3 sp_1.5-1 plotly_4.10.1
[112] spatstat.sparse_3.0-0 xml2_1.3.3 bslib_0.4.1
[115] XVector_0.36.0 rvest_1.0.3 digest_0.6.30
[118] sctransform_0.3.5 RcppAnnoy_0.0.20 spatstat.data_3.0-0
[121] rmarkdown_2.18 cellranger_1.1.0 leiden_0.4.3
[124] uwot_0.1.14 shiny_1.7.3 lifecycle_1.0.3
[127] nlme_3.1-160 jsonlite_1.8.3 carData_3.0-5
[130] fansi_1.0.3 pillar_1.8.1 lattice_0.20-45
[133] fastmap_1.1.0 httr_1.4.4 survival_3.4-0
[136] glue_1.6.2 png_0.1-8 bit_4.0.5
[139] stringi_1.7.8 sass_0.4.4 irlba_2.3.5.1
[142] future.apply_1.10.0